
 -
-
-
-
-
-
[📚Documentation](https://deeplabcut.github.io/DeepLabCut/README.html) |
[🛠️ Installation](https://deeplabcut.github.io/DeepLabCut/docs/installation.html) |
[🌎 Home Page](https://www.deeplabcut.org) |
@@ -57,7 +51,7 @@
**DeepLabCut™️** is a toolbox for state-of-the-art markerless pose estimation of animals performing various behaviors. As long as you can see (label) what you want to track, you can use this toolbox, as it is animal and object agnostic. [Read a short development and application summary below](https://github.com/DeepLabCut/DeepLabCut#why-use-deeplabcut).
-# [Installation: how to install DeepLabCut](https://deeplabcut.github.io/DeepLabCut/docs/installation.html)
+# [Installation](https://deeplabcut.github.io/DeepLabCut/docs/installation.html)
Please click the link above for all the information you need to get started! Please note that currently we support only Python 3.10+ (see conda files for guidance).
@@ -80,39 +74,47 @@ pip install --pre "deeplabcut[gui]"
or `pip install --pre "deeplabcut"` (headless
version with PyTorch)!
-To use the TensorFlow (TF) engine (requires Python 3.10; TF up to v2.10 supported on Windows,
-up to v2.12 on other platforms): you'll need to run `pip install "deeplabcut[gui,tf]"`
-(which includes all functions plus GUIs) or `pip install "deeplabcut[tf]"` (headless
-version with PyTorch and TensorFlow). We aim to depreciate the TF part in 2027.
+To use the TensorFlow (TF) engine: you'll need to run `pip install "deeplabcut[gui,tf]"` or `pip install "deeplabcut[tf]"` (headless version with TF).
+We aim to deprecate the tensorflow backend in version 3.2 (release date TBD).
We recommend using our conda file, see [here](https://github.com/DeepLabCut/DeepLabCut/blob/main/conda-environments/README.md) or the [`deeplabcut-docker` package](https://github.com/DeepLabCut/DeepLabCut/tree/main/docker).
-# [Documentation: The DeepLabCut Process](https://deeplabcut.github.io/DeepLabCut/README.html)
+
+# Documentation: The DeepLabCut Process
Our docs walk you through using DeepLabCut, and key API points. For an overview of the toolbox and workflow for project management, see our step-by-step at [Nature Protocols paper](https://doi.org/10.1038/s41596-019-0176-0).
-For a deeper understanding and more resources for you to get started with Python and DeepLabCut, please check out our free online course! https://deeplabcut.github.io/DeepLabCut/docs/course.html
+
+
-
-
-
-
-
-
[📚Documentation](https://deeplabcut.github.io/DeepLabCut/README.html) |
[🛠️ Installation](https://deeplabcut.github.io/DeepLabCut/docs/installation.html) |
[🌎 Home Page](https://www.deeplabcut.org) |
@@ -57,7 +51,7 @@
**DeepLabCut™️** is a toolbox for state-of-the-art markerless pose estimation of animals performing various behaviors. As long as you can see (label) what you want to track, you can use this toolbox, as it is animal and object agnostic. [Read a short development and application summary below](https://github.com/DeepLabCut/DeepLabCut#why-use-deeplabcut).
-# [Installation: how to install DeepLabCut](https://deeplabcut.github.io/DeepLabCut/docs/installation.html)
+# [Installation](https://deeplabcut.github.io/DeepLabCut/docs/installation.html)
Please click the link above for all the information you need to get started! Please note that currently we support only Python 3.10+ (see conda files for guidance).
@@ -80,39 +74,47 @@ pip install --pre "deeplabcut[gui]"
or `pip install --pre "deeplabcut"` (headless
version with PyTorch)!
-To use the TensorFlow (TF) engine (requires Python 3.10; TF up to v2.10 supported on Windows,
-up to v2.12 on other platforms): you'll need to run `pip install "deeplabcut[gui,tf]"`
-(which includes all functions plus GUIs) or `pip install "deeplabcut[tf]"` (headless
-version with PyTorch and TensorFlow). We aim to depreciate the TF part in 2027.
+To use the TensorFlow (TF) engine: you'll need to run `pip install "deeplabcut[gui,tf]"` or `pip install "deeplabcut[tf]"` (headless version with TF).
+We aim to deprecate the tensorflow backend in version 3.2 (release date TBD).
We recommend using our conda file, see [here](https://github.com/DeepLabCut/DeepLabCut/blob/main/conda-environments/README.md) or the [`deeplabcut-docker` package](https://github.com/DeepLabCut/DeepLabCut/tree/main/docker).
-# [Documentation: The DeepLabCut Process](https://deeplabcut.github.io/DeepLabCut/README.html)
+
+# Documentation: The DeepLabCut Process
Our docs walk you through using DeepLabCut, and key API points. For an overview of the toolbox and workflow for project management, see our step-by-step at [Nature Protocols paper](https://doi.org/10.1038/s41596-019-0176-0).
-For a deeper understanding and more resources for you to get started with Python and DeepLabCut, please check out our free online course! https://deeplabcut.github.io/DeepLabCut/docs/course.html
+
+

@@ -145,49 +152,51 @@ We currently provide state-of-the-art performance for animal pose estimation and **Left:** Due to transfer learning it requires **little training data** for multiple, challenging behaviors (see [Mathis et al. 2018](https://www.nature.com/articles/s41593-018-0209-y) for details). **Mid Left:** The feature detectors are robust to video compression (see [Mathis/Warren](https://www.biorxiv.org/content/early/2018/10/30/457242) for details). **Mid Right:** It allows 3D pose estimation with a single network and camera (see [Mathis/Warren](https://www.biorxiv.org/content/early/2018/10/30/457242)). **Right:** It allows 3D pose estimation with a single network trained on data from multiple cameras together with standard triangulation methods (see [Nath* and Mathis* et al. 2019](https://doi.org/10.1038/s41596-019-0176-0)). -**DeepLabCut** is embedding in a larger open-source eco-system, providing behavioral tracking for neuroscience, ecology, medical, and technical applications. Moreover, many new tools are being actively developed. See [DLC-Utils](https://github.com/DeepLabCut/DLCutils) for some helper code. +### Ecosystem + +**DeepLabCut** is part of a larger open-source eco-system, providing behavioral tracking for neuroscience, ecology, medical, and technical applications. +Moreover, many new tools are being actively developed. See [DLC-Utils](https://github.com/DeepLabCut/DLCutils) for some helper code.

+ +
+
+  +
+


- -
-
-  -
-
@@ -87,85 +100,134 @@ This page contains a list of the essential functions of DeepLabCut as well as de
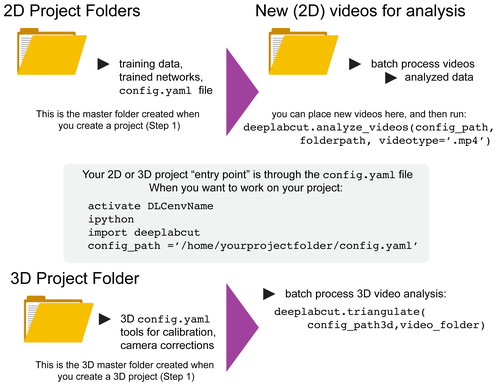
-You can have as many projects on your computer as you wish. You can have DeepLabCut installed in an [environment](../conda-environments/README.md) and always exit and return to this environment to run the code. You just need to point to the correct ``config.yaml`` file to [jump back in](/docs/UseOverviewGuide.md#tips-for-daily-use)! The documentation below will take you through the individual steps. +You can have as many projects on your computer as you wish. +You can have DeepLabCut installed in a {ref}`conda environment
 +## Network training
+
+### Creating a training dataset
Before training your model, the first step is to assemble your training dataset.
+This involves:
-**Create Training Dataset:** Move to the corresponding tab and click **`Create Training Dataset`**. For starters, the default settings will do just fine. While there are more powerful models and data augmentations you might want to consider, you can trust that for most projects the defaults are an ideal place to start.
+- Splitting labeled data into training and evaluation subsets
+- Creating each shuffle folder with the model configuration ready for training.
-> 💡 **Note:** This guide assumes you have a GPU on your local machine. If you're CPU-bound and finding training challenging, consider using Google Colab. Our [Colab Guide](https://colab.research.google.com/github/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb) can help you get started!
+**Create Training Dataset:** Move to the corresponding tab and click **`Create Training Dataset`**. For starters, the default settings will do just fine. While there are more powerful models and data augmentations you might want to consider, you can trust that for most projects the defaults are a good place to start.
-## Kickstarting the Training Process
+```{note}
+This guide assumes you have a (CUDA-enabled) GPU on your local machine. If you're CPU-bound and training is not feasible, consider using Google Colab. Our [Colab Guide](https://colab.research.google.com/github/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb) can help you get started!
+```
+
+### Starting the training process
With your training dataset ready, it's time to train your model.
@@ -33,31 +46,34 @@ You can keep an eye on the training progress via your terminal window. This will

-## Evaluate the Network
+## Network evaluation
After training, it's time to see how well your model performs.
-### Steps to Evaluate the Network
+### Step-by-step
1. Find and click on the **`Evaluate Network`** tab.
-2. **Choose Evaluation Options:**
+1. **Choose Evaluation Options:**
- **Plot Predictions:** Select this to visualize the model's predictions, similar to standard DeepLabCut (DLC) evaluations.
- **Compare Bodyparts:** Opt to compare all the bodyparts for a comprehensive evaluation.
-3. Click the **`Evaluate Network`** button, located on the right side of the main window.
-
->💡 Tip: If you wish to evaluate all saved snapshots, go to the configuration file and change the `snapshotindex` parameter to `all`.
+1. Click the **`Evaluate Network`** button, located on the right side of the main window.
+```{tip}
+If you wish to evaluate all saved snapshots, go to the configuration file and change the `snapshotindex` parameter to `all`.
+```
-### Understanding the Evaluation Results
+### Interpreting the results
- **Performance Metrics:** DLC will assess the latest snapshot of your model, generating a `.CSV` file with performance
-metrics. This file is stored in the **`evaluation-results`** (for TensorFlow models) or the
-**`evaluation-results-pytorch`** (for PyTorch models) folder within your project.
+ metrics. This file is stored in the **`evaluation-results`** (for TensorFlow models) or the
+ **`evaluation-results-pytorch`** (for PyTorch models) folder within your project.
+
-)
- **Visual Feedback:** Additionally, DLC creates subfolders containing your frames overlaid with both the labeled bodyparts and the model's predictions, allowing you to visually gauge the network's performance.
-)
+
+
+## Next steps
-## Next, head over the beginner guide for [using your new neural network for video analysis](video-analysis)
+Head over the {ref}`file:video-analysis-gui` section to learn about applying your trained model to videos, and creating labeled videos with the results of your analysis!
diff --git a/docs/beginner-guides/beginners-guide.md b/docs/beginner-guides/beginners-guide.md
index a4df28db5..43bce7f1c 100644
--- a/docs/beginner-guides/beginners-guide.md
+++ b/docs/beginner-guides/beginners-guide.md
@@ -5,24 +5,33 @@ deeplabcut:
ignore: false
visibility: online
status: outdated
- recommendation: update
- notes: "While it could seem like a useful page for beginners, duplicating installation instructions is not ideal for maintenance. This is also mixing installation/setup with a GUI guide, which should be in its own section/page. This puts into question the reason of existence of this page, as it would end up being two links to different sections. I would rather have well-made, accurate installation and GUI guides, and if there are beginner-relevant information that really cannot fit into those, then we can have a 'beginner's guide' that links to those and has the extra info. I would suggest reviewing whether this style of docs should remain at all, but if we want to keep them revising the approach may be needed."
+ recommendation: move
+ notes: Move to GUI section.
---
-(beginners-guide)=
-# Using DeepLabCut
+
+(file:beginners-guide)=
+
+# Using the DeepLabCut GUI
+
+## Network training
+
+### Creating a training dataset
Before training your model, the first step is to assemble your training dataset.
+This involves:
-**Create Training Dataset:** Move to the corresponding tab and click **`Create Training Dataset`**. For starters, the default settings will do just fine. While there are more powerful models and data augmentations you might want to consider, you can trust that for most projects the defaults are an ideal place to start.
+- Splitting labeled data into training and evaluation subsets
+- Creating each shuffle folder with the model configuration ready for training.
-> 💡 **Note:** This guide assumes you have a GPU on your local machine. If you're CPU-bound and finding training challenging, consider using Google Colab. Our [Colab Guide](https://colab.research.google.com/github/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb) can help you get started!
+**Create Training Dataset:** Move to the corresponding tab and click **`Create Training Dataset`**. For starters, the default settings will do just fine. While there are more powerful models and data augmentations you might want to consider, you can trust that for most projects the defaults are a good place to start.
-## Kickstarting the Training Process
+```{note}
+This guide assumes you have a (CUDA-enabled) GPU on your local machine. If you're CPU-bound and training is not feasible, consider using Google Colab. Our [Colab Guide](https://colab.research.google.com/github/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb) can help you get started!
+```
+
+### Starting the training process
With your training dataset ready, it's time to train your model.
@@ -33,31 +46,34 @@ You can keep an eye on the training progress via your terminal window. This will

-## Evaluate the Network
+## Network evaluation
After training, it's time to see how well your model performs.
-### Steps to Evaluate the Network
+### Step-by-step
1. Find and click on the **`Evaluate Network`** tab.
-2. **Choose Evaluation Options:**
+1. **Choose Evaluation Options:**
- **Plot Predictions:** Select this to visualize the model's predictions, similar to standard DeepLabCut (DLC) evaluations.
- **Compare Bodyparts:** Opt to compare all the bodyparts for a comprehensive evaluation.
-3. Click the **`Evaluate Network`** button, located on the right side of the main window.
-
->💡 Tip: If you wish to evaluate all saved snapshots, go to the configuration file and change the `snapshotindex` parameter to `all`.
+1. Click the **`Evaluate Network`** button, located on the right side of the main window.
+```{tip}
+If you wish to evaluate all saved snapshots, go to the configuration file and change the `snapshotindex` parameter to `all`.
+```
-### Understanding the Evaluation Results
+### Interpreting the results
- **Performance Metrics:** DLC will assess the latest snapshot of your model, generating a `.CSV` file with performance
-metrics. This file is stored in the **`evaluation-results`** (for TensorFlow models) or the
-**`evaluation-results-pytorch`** (for PyTorch models) folder within your project.
+ metrics. This file is stored in the **`evaluation-results`** (for TensorFlow models) or the
+ **`evaluation-results-pytorch`** (for PyTorch models) folder within your project.
+
-)
- **Visual Feedback:** Additionally, DLC creates subfolders containing your frames overlaid with both the labeled bodyparts and the model's predictions, allowing you to visually gauge the network's performance.
-)
+
+
+## Next steps
-## Next, head over the beginner guide for [using your new neural network for video analysis](video-analysis)
+Head over the {ref}`file:video-analysis-gui` section to learn about applying your trained model to videos, and creating labeled videos with the results of your analysis!
diff --git a/docs/beginner-guides/beginners-guide.md b/docs/beginner-guides/beginners-guide.md
index a4df28db5..43bce7f1c 100644
--- a/docs/beginner-guides/beginners-guide.md
+++ b/docs/beginner-guides/beginners-guide.md
@@ -5,24 +5,33 @@ deeplabcut:
ignore: false
visibility: online
status: outdated
- recommendation: update
- notes: "While it could seem like a useful page for beginners, duplicating installation instructions is not ideal for maintenance. This is also mixing installation/setup with a GUI guide, which should be in its own section/page. This puts into question the reason of existence of this page, as it would end up being two links to different sections. I would rather have well-made, accurate installation and GUI guides, and if there are beginner-relevant information that really cannot fit into those, then we can have a 'beginner's guide' that links to those and has the extra info. I would suggest reviewing whether this style of docs should remain at all, but if we want to keep them revising the approach may be needed."
+ recommendation: move
+ notes: Move to GUI section.
---
-(beginners-guide)=
-# Using DeepLabCut
+
+(file:beginners-guide)=
+
+# Using the DeepLabCut GUI
+
 This guide, and related pages, are meant as a very-new-to-python beginner guide to DeepLabCut. After you are comfortable with this material we recommend then jumping into the more detailed User Guides!
-- **ProTip:** For even more 'in-depth' understanding, head over to check out the [DeepLabCut Course](https://deeplabcut.github.io/DeepLabCut/docs/course.html), which provides a deeper dive into the science behind DeepLabCut.
+
+
+
## Installation
Before you begin, make sure that DeepLabCut is installed on your system.
-- **ProTip:** For detailed installation instructions, geared towards a bit more advanced users, refer to the [Full Installation Guide](https://deeplabcut.github.io/DeepLabCut/docs/installation.html).
+Please see the {ref}`installation page
This guide, and related pages, are meant as a very-new-to-python beginner guide to DeepLabCut. After you are comfortable with this material we recommend then jumping into the more detailed User Guides!
-- **ProTip:** For even more 'in-depth' understanding, head over to check out the [DeepLabCut Course](https://deeplabcut.github.io/DeepLabCut/docs/course.html), which provides a deeper dive into the science behind DeepLabCut.
+
+
+
## Installation
Before you begin, make sure that DeepLabCut is installed on your system.
-- **ProTip:** For detailed installation instructions, geared towards a bit more advanced users, refer to the [Full Installation Guide](https://deeplabcut.github.io/DeepLabCut/docs/installation.html).
+Please see the {ref}`installation page -**Edit the Configuration File**
+The configuration file (`config.yaml`) is the central record of files in your project, as well as the settings for your models.
+As a YAML file, it can be edited manually, but the GUI provides an easy way to edit it without needing to know the YAML format. In this guide, we will show you how to edit the configuration file using the GUI.
-After creating your DeepLabCut project, you'll go to the main GUI window, where you'll start managing your project from the Project Management Tab.
+## Editing the configuration
-**Accessing the Configuration File**
+After creating your DeepLabCut project, you'll be shown the main GUI window, where you can manage your project from the Project Management Tab.
- **Locate the Configuration File:** At the top of the main window, you'll find the file path to the configuration file.
-- **Edit the File:** Click on **`Edit config.yaml`**. This action allows you to:
- - Define the bodyparts you wish to track.
- - Outline the skeleton structure (optional!).
+- **Edit the File:** Click on **`Edit config.yaml`**.
+ - A **`Configuration Editor`** window will open, displaying all the configuration details.
+ - You will need to modify some of these settings to align with your experiment.
+ - For example:
+ - Update or define the bodyparts you wish to track.
+ - *Optional:* Outline the skeleton structure.
-A **`Configuration Editor`** window will open, displaying all the configuration details. You'll need to modify some of these settings to align with your research requirements.
+## Step-by-step configuration walkthrough
-## Steps to Edit the Configuration
-
-### 1. Defining Bodyparts
+### Defining & updating bodyparts
- **Locate the Bodyparts Section:** In the Configuration Editor, find the **`bodyparts`** category.
- **Modify the List:** Click on the arrow next to **`bodyparts`** to expand the list. Here, you can:
- Update the list with the names of the bodyparts relevant to your study.
- Add more entries by right-clicking on a row number and selecting **`Insert`**.
-

+
-### 2. Defining the Skeleton
+### Defining the skeleton
- **Navigate to the Skeleton Section:** Scroll down to the **`skeleton`** category.
-- **Adjust the Skeleton List:** Click on the arrow to expand this section. You can then:
- - Update the pairs of bodyparts to define the skeleton structure of your model.
+- **Adjust the Skeleton List:** Click on the arrow to expand this section.
+ - You can then update the list of bodypart pairs: i.e. the connections that define the skeleton structure of your model.
+ - In the list of bodypart pairs, each pair has an index. (ranging from 0 to the total number of pairs in the skeleton).
+ - Each item of the pair (also indexed; 0 or 1) has a value: the name of the bodypart.
+ - Each pair of two bodyparts represents a connection, where all connections together make the skeleton.

-> 💡 **Tip:** If you're new to DeepLabCut, spend some time visualizing how the chosen bodyparts can be connected effectively to form a coherent skeleton.
+```{tip}
+Spend some time visualizing how the chosen bodyparts can be connected effectively to form a coherent, visually helpful skeleton.
+```
-### Saving Your Changes
+### Saving changes
- **Save the Configuration:** Once you're satisfied with the modifications, click **`Save`**. This will store your changes and return you to the main GUI window.
-## Next, head over the beginner guide for [Labeling your data](labeling)
+## Next steps
+
+Head over the guide for the {ref}`file:labeling-gui`, which will show you how to label your data using the napari-based labeling GUI.
diff --git a/docs/beginner-guides/video-analysis.md b/docs/beginner-guides/video-analysis.md
index 5ed589253..ac667881b 100644
--- a/docs/beginner-guides/video-analysis.md
+++ b/docs/beginner-guides/video-analysis.md
@@ -6,23 +6,30 @@ deeplabcut:
visibility: online
status: viable
recommendation: move
- notes: "As mentioned on oher beginner-guides/ docs, this should be part of the GUI section."
+ notes: As mentioned on oher beginner-guides/ docs, this should be part of the GUI section.
---
-# Video Analysis with DeepLabCut
-
-**Edit the Configuration File**
+The configuration file (`config.yaml`) is the central record of files in your project, as well as the settings for your models.
+As a YAML file, it can be edited manually, but the GUI provides an easy way to edit it without needing to know the YAML format. In this guide, we will show you how to edit the configuration file using the GUI.
-After creating your DeepLabCut project, you'll go to the main GUI window, where you'll start managing your project from the Project Management Tab.
+## Editing the configuration
-**Accessing the Configuration File**
+After creating your DeepLabCut project, you'll be shown the main GUI window, where you can manage your project from the Project Management Tab.
- **Locate the Configuration File:** At the top of the main window, you'll find the file path to the configuration file.
-- **Edit the File:** Click on **`Edit config.yaml`**. This action allows you to:
- - Define the bodyparts you wish to track.
- - Outline the skeleton structure (optional!).
+- **Edit the File:** Click on **`Edit config.yaml`**.
+ - A **`Configuration Editor`** window will open, displaying all the configuration details.
+ - You will need to modify some of these settings to align with your experiment.
+ - For example:
+ - Update or define the bodyparts you wish to track.
+ - *Optional:* Outline the skeleton structure.
-A **`Configuration Editor`** window will open, displaying all the configuration details. You'll need to modify some of these settings to align with your research requirements.
+## Step-by-step configuration walkthrough
-## Steps to Edit the Configuration
-
-### 1. Defining Bodyparts
+### Defining & updating bodyparts
- **Locate the Bodyparts Section:** In the Configuration Editor, find the **`bodyparts`** category.
- **Modify the List:** Click on the arrow next to **`bodyparts`** to expand the list. Here, you can:
- Update the list with the names of the bodyparts relevant to your study.
- Add more entries by right-clicking on a row number and selecting **`Insert`**.
-

+
-### 2. Defining the Skeleton
+### Defining the skeleton
- **Navigate to the Skeleton Section:** Scroll down to the **`skeleton`** category.
-- **Adjust the Skeleton List:** Click on the arrow to expand this section. You can then:
- - Update the pairs of bodyparts to define the skeleton structure of your model.
+- **Adjust the Skeleton List:** Click on the arrow to expand this section.
+ - You can then update the list of bodypart pairs: i.e. the connections that define the skeleton structure of your model.
+ - In the list of bodypart pairs, each pair has an index. (ranging from 0 to the total number of pairs in the skeleton).
+ - Each item of the pair (also indexed; 0 or 1) has a value: the name of the bodypart.
+ - Each pair of two bodyparts represents a connection, where all connections together make the skeleton.

-> 💡 **Tip:** If you're new to DeepLabCut, spend some time visualizing how the chosen bodyparts can be connected effectively to form a coherent skeleton.
+```{tip}
+Spend some time visualizing how the chosen bodyparts can be connected effectively to form a coherent, visually helpful skeleton.
+```
-### Saving Your Changes
+### Saving changes
- **Save the Configuration:** Once you're satisfied with the modifications, click **`Save`**. This will store your changes and return you to the main GUI window.
-## Next, head over the beginner guide for [Labeling your data](labeling)
+## Next steps
+
+Head over the guide for the {ref}`file:labeling-gui`, which will show you how to label your data using the napari-based labeling GUI.
diff --git a/docs/beginner-guides/video-analysis.md b/docs/beginner-guides/video-analysis.md
index 5ed589253..ac667881b 100644
--- a/docs/beginner-guides/video-analysis.md
+++ b/docs/beginner-guides/video-analysis.md
@@ -6,23 +6,30 @@ deeplabcut:
visibility: online
status: viable
recommendation: move
- notes: "As mentioned on oher beginner-guides/ docs, this should be part of the GUI section."
+ notes: As mentioned on oher beginner-guides/ docs, this should be part of the GUI section.
---
-# Video Analysis with DeepLabCut
- +(file:video-analysis-gui)=
+
+# Video analysis in the GUI
+
+
+(file:video-analysis-gui)=
+
+# Video analysis in the GUI
+
+ After training and evaluating your model, the next step is to apply it to your videos.
-**How to Analyze Videos**
+## Analyzing videos with your trained model
+
+### Step-by-step
1. **Navigate to the 'Analyze Videos' Tab:** Begin applying your trained model to video data here.
-2. **Select Your Video Format and Files:**
- - **Choose Video Format:** Pick the format of your video (`.mp4`, `.avi`, `.mkv`, or `.mov`).
- - **Select Videos:** Click **`Select Videos`** to find and open your video file.
+1. **Select Your Video Format and Files:**
+
+- **Choose Video Format:** Pick the format of your video (`.mp4`, `.avi`, `.mkv`, or `.mov`).
+- **Select Videos:** Click **`Select Videos`** to find and open your video file.
+
3. **Start Analysis:** Click **`Analyze`**. The analysis time depends on video length and resolution. Track progress in the terminal or Anaconda prompt.
-## Reviewing Analysis Results
+### Reviewing analysis results
- **Find Results in Your Project Folder:** After analysis, go to your project's video folder.
- **Analysis Files:** Look also for a `.metapickle`, an `.h5`, and possibly a `.csv` file for detailed analysis data.
@@ -30,16 +37,21 @@ After training and evaluating your model, the next step is to apply it to your v

-## Creating a Labeled Video
+## Generating labeled videos
+
+### Create a labeled video
1. **Go to 'Create Labeled Video' Tab:** The previously analyzed video should be selected.
-2. If not already selected, choose your video.
-3. Click **`Create Videos`**.
+1. If not already selected, choose your video.
+1. Click **`Create Videos`**.
-## Viewing the Labeled Video
+### View the labeled video
- Your labeled video will be in your video folder, named after the original video plus model details and 'labeled'.
-- Watch the video to assess the model's labeling accuracy.
+- Use it in your results, or perform downstream analyses with it!
+
+## Next steps
+
+
-## Happy DeepLabCutting!
-- Check out the more advanced user guides for even more options!
+Check our more advanced guides, and consider reading more about models, augmentations and other parameters to further optimize your model and analysis!
diff --git a/docs/course.md b/docs/course.md
index a47610c78..94d4c24d0 100644
--- a/docs/course.md
+++ b/docs/course.md
@@ -7,16 +7,17 @@ deeplabcut:
status: outdated
recommendation: archive
---
+
# DeepLabCut Self-paced Course
-::::{warning}
+::::\{warning}
This course was designed for DLC 2.
An updated version for DLC 3 is in the works.
::::
Do you have video of animal behaviors? Step 1: Get Poses ...
-
After training and evaluating your model, the next step is to apply it to your videos.
-**How to Analyze Videos**
+## Analyzing videos with your trained model
+
+### Step-by-step
1. **Navigate to the 'Analyze Videos' Tab:** Begin applying your trained model to video data here.
-2. **Select Your Video Format and Files:**
- - **Choose Video Format:** Pick the format of your video (`.mp4`, `.avi`, `.mkv`, or `.mov`).
- - **Select Videos:** Click **`Select Videos`** to find and open your video file.
+1. **Select Your Video Format and Files:**
+
+- **Choose Video Format:** Pick the format of your video (`.mp4`, `.avi`, `.mkv`, or `.mov`).
+- **Select Videos:** Click **`Select Videos`** to find and open your video file.
+
3. **Start Analysis:** Click **`Analyze`**. The analysis time depends on video length and resolution. Track progress in the terminal or Anaconda prompt.
-## Reviewing Analysis Results
+### Reviewing analysis results
- **Find Results in Your Project Folder:** After analysis, go to your project's video folder.
- **Analysis Files:** Look also for a `.metapickle`, an `.h5`, and possibly a `.csv` file for detailed analysis data.
@@ -30,16 +37,21 @@ After training and evaluating your model, the next step is to apply it to your v

-## Creating a Labeled Video
+## Generating labeled videos
+
+### Create a labeled video
1. **Go to 'Create Labeled Video' Tab:** The previously analyzed video should be selected.
-2. If not already selected, choose your video.
-3. Click **`Create Videos`**.
+1. If not already selected, choose your video.
+1. Click **`Create Videos`**.
-## Viewing the Labeled Video
+### View the labeled video
- Your labeled video will be in your video folder, named after the original video plus model details and 'labeled'.
-- Watch the video to assess the model's labeling accuracy.
+- Use it in your results, or perform downstream analyses with it!
+
+## Next steps
+
+
-## Happy DeepLabCutting!
-- Check out the more advanced user guides for even more options!
+Check our more advanced guides, and consider reading more about models, augmentations and other parameters to further optimize your model and analysis!
diff --git a/docs/course.md b/docs/course.md
index a47610c78..94d4c24d0 100644
--- a/docs/course.md
+++ b/docs/course.md
@@ -7,16 +7,17 @@ deeplabcut:
status: outdated
recommendation: archive
---
+
# DeepLabCut Self-paced Course
-::::{warning}
+::::\{warning}
This course was designed for DLC 2.
An updated version for DLC 3 is in the works.
::::
Do you have video of animal behaviors? Step 1: Get Poses ...
-  +
+ This document is an outline of resources for a course for those wanting to learn to use `Python` and `DeepLabCut`.
We expect it to take *roughly* 1-2 weeks to get through if you do it rigorously. To get the basics, it should take 1-2 days.
@@ -27,14 +28,13 @@ We expect it to take *roughly* 1-2 weeks to get through if you do it rigorously.
This document is an outline of resources for a course for those wanting to learn to use `Python` and `DeepLabCut`.
We expect it to take *roughly* 1-2 weeks to get through if you do it rigorously. To get the basics, it should take 1-2 days.
@@ -27,14 +28,13 @@ We expect it to take *roughly* 1-2 weeks to get through if you do it rigorously.
 -
## Installation:
You need Python and DeepLabCut installed!
-- [See these "beginner docs" for help!](beginners-guide)
-- **WATCH:** overview of conda: [Python Tutorial: Anaconda - Installation and Using Conda](https://www.youtube.com/watch?v=YJC6ldI3hWk)
+- \[See these "beginner docs" for help!\](file:beginners-guide)
+- **WATCH:** overview of conda: [Python Tutorial: Anaconda - Installation and Using Conda](https://www.youtube.com/watch?v=YJC6ldI3hWk)
## Outline:
@@ -47,93 +47,91 @@ You need Python and DeepLabCut installed!
- **Learning:** learning and teaching signal processing, and overview from Prof. Demba Ba [talk at JupyterCon](https://www.youtube.com/watch?v=ywz-LLYwkQQ)
- **DEMO:** Can I DEMO DEEPLABCUT (DLC) quickly?
- - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
- - AND follow along with me: [Video Tutorial!](https://www.youtube.com/watch?v=DRT-Cq2vdWs)
+ - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
+ - AND follow along with me: [Video Tutorial!](https://www.youtube.com/watch?v=DRT-Cq2vdWs)
- **WATCH:** How do you know DLC is installed properly? (i.e. how to use our test script!) [Video Tutorial!](https://youtu.be/IOWtKn3l33s)
-
-
## Installation:
You need Python and DeepLabCut installed!
-- [See these "beginner docs" for help!](beginners-guide)
-- **WATCH:** overview of conda: [Python Tutorial: Anaconda - Installation and Using Conda](https://www.youtube.com/watch?v=YJC6ldI3hWk)
+- \[See these "beginner docs" for help!\](file:beginners-guide)
+- **WATCH:** overview of conda: [Python Tutorial: Anaconda - Installation and Using Conda](https://www.youtube.com/watch?v=YJC6ldI3hWk)
## Outline:
@@ -47,93 +47,91 @@ You need Python and DeepLabCut installed!
- **Learning:** learning and teaching signal processing, and overview from Prof. Demba Ba [talk at JupyterCon](https://www.youtube.com/watch?v=ywz-LLYwkQQ)
- **DEMO:** Can I DEMO DEEPLABCUT (DLC) quickly?
- - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
- - AND follow along with me: [Video Tutorial!](https://www.youtube.com/watch?v=DRT-Cq2vdWs)
+ - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
+ - AND follow along with me: [Video Tutorial!](https://www.youtube.com/watch?v=DRT-Cq2vdWs)
- **WATCH:** How do you know DLC is installed properly? (i.e. how to use our test script!) [Video Tutorial!](https://youtu.be/IOWtKn3l33s)
-
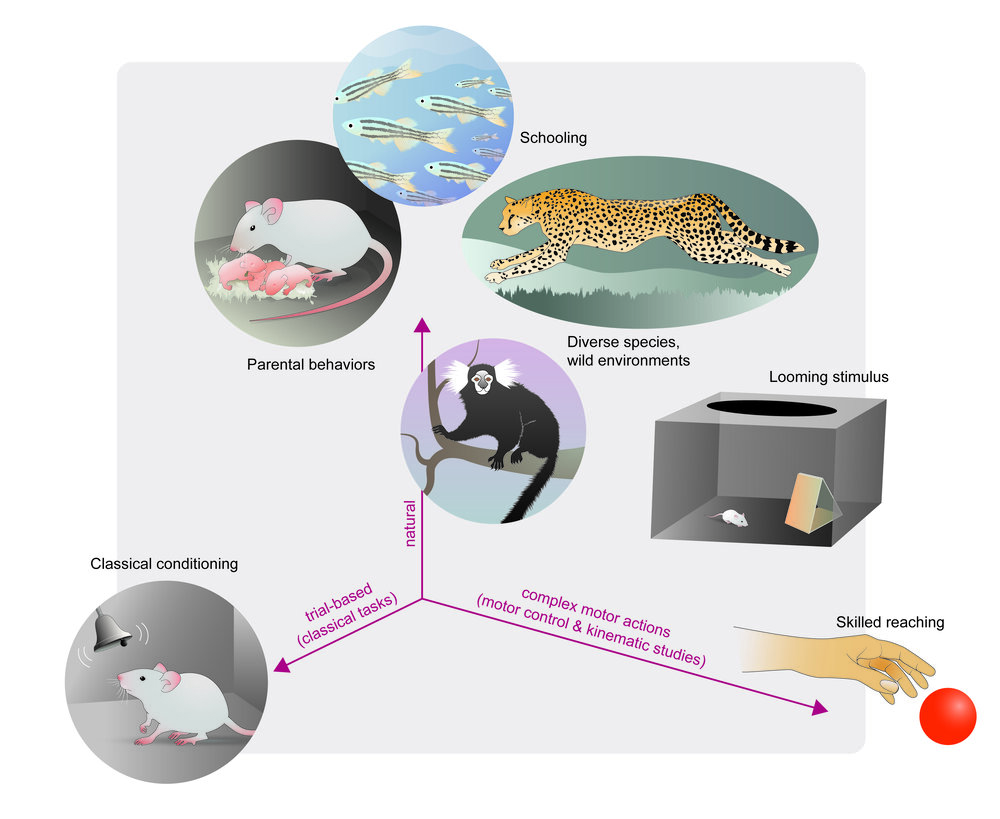 - **REVIEW PAPER:** The state of animal pose estimation w/ deep learning i.e. "Deep learning tools for the measurement of animal behavior in neuroscience" [arXiv](https://arxiv.org/abs/1909.13868) & [published version](https://www.sciencedirect.com/science/article/pii/S0959438819301151)
- **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
-
- **WATCH:** There are a lot of docs... where to begin: [Video Tutorial!](https://www.youtube.com/watch?v=A9qZidI7tL8)
### **Module 1: getting started on data**
**What you need:** any videos where you can see the animals/objects, etc.
You can use our demo videos, grab some from the internet, or use whatever older data you have. Any camera, color/monochrome, etc will work. Find diverse videos, and label what you want to track well :)
-- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo 😊
- - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part1-labeling.pdf)
- - **READ ME PLEASE:** [DeepLabCut, the science](https://rdcu.be/4Rep)
- - **READ ME PLEASE:** [DeepLabCut, the user guide](https://rdcu.be/bHpHN)
- - **WATCH:** Video tutorial 1: [using the Project Manager GUI](https://www.youtube.com/watch?v=KcXogR-p5Ak)
- - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
- - **WATCH:** Video tutorial 2: [using the Project Manager GUI for multi-animal pose estimation](https://www.youtube.com/watch?v=Kp-stcTm77g)
- - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
- - **WATCH:** Video tutorial 3: [using ipython/pythonw (more functions!)](https://www.youtube.com/watch?v=7xwOhUcIGio)
- - multi-animal DLC: [labeling](https://www.youtube.com/watch?v=Kp-stcTm77g)
- - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
+- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo 😊
+ - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part1-labeling.pdf)
+ - **READ ME PLEASE:** [DeepLabCut, the science](https://rdcu.be/4Rep)
+ - **READ ME PLEASE:** [DeepLabCut, the user guide](https://rdcu.be/bHpHN)
+ - **WATCH:** Video tutorial 1: [using the Project Manager GUI](https://www.youtube.com/watch?v=KcXogR-p5Ak)
+ - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
+ - **WATCH:** Video tutorial 2: [using the Project Manager GUI for multi-animal pose estimation](https://www.youtube.com/watch?v=Kp-stcTm77g)
+ - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
+ - **WATCH:** Video tutorial 3: [using ipython/pythonw (more functions!)](https://www.youtube.com/watch?v=7xwOhUcIGio)
+ - multi-animal DLC: [labeling](https://www.youtube.com/watch?v=Kp-stcTm77g)
+ - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
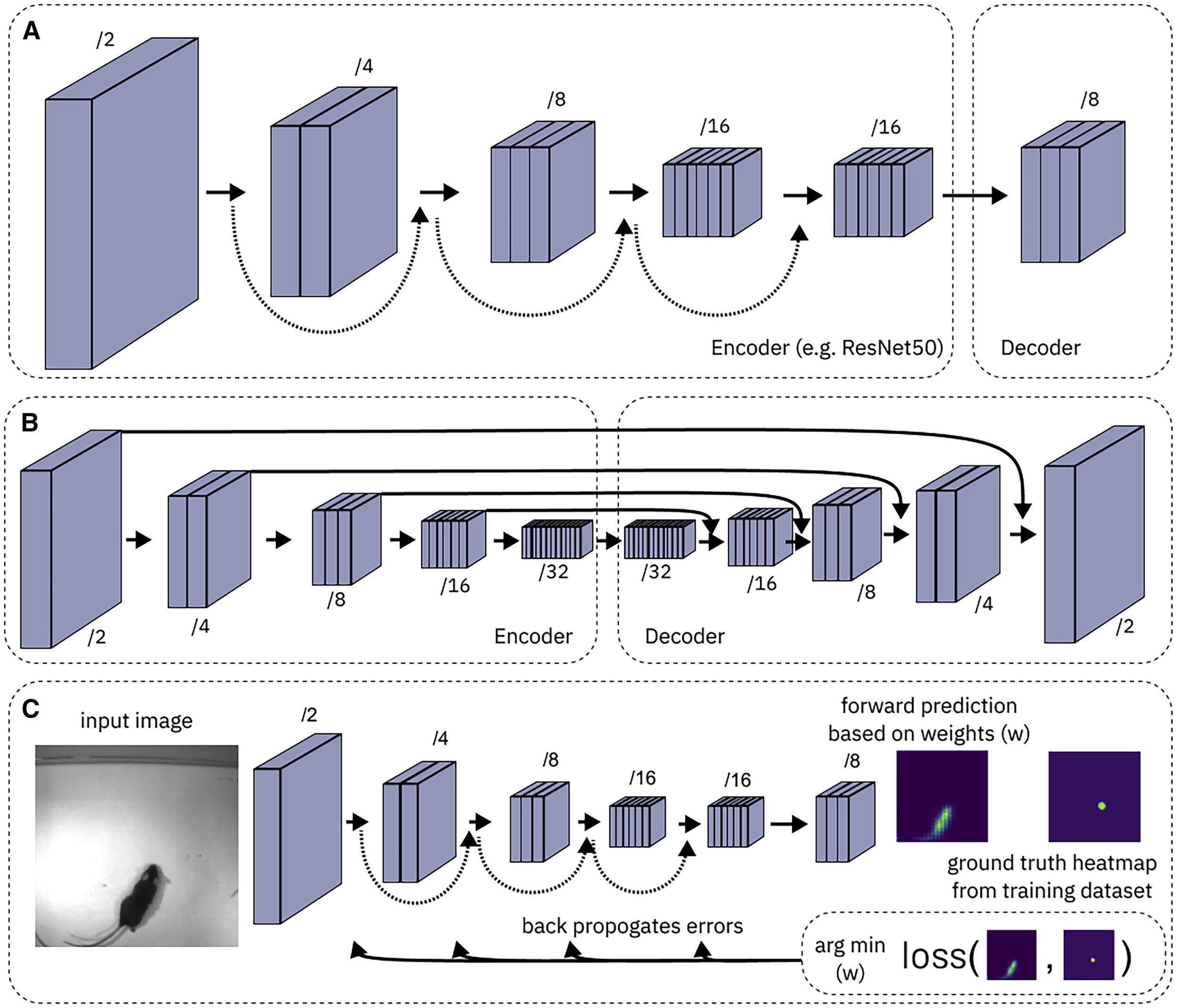
### **Module 2: Neural Networks**
- - **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part2-network.pdf)
- - **READ ME PLEASE:** [What are convolutional neural networks?](https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
+- **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part2-network.pdf)
+
+- **READ ME PLEASE:** [What are convolutional neural networks?](https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
- - **READ ME PLEASE:** Here is a new paper from us describing challenges in robust pose estimation, why PRE-TRAINING really matters - which was our major scientific contribution to low-data input pose-estimation - and it describes new networks that are available to you. [Pretraining boosts out-of-domain robustness for pose estimation](https://paperswithcode.com/paper/pretraining-boosts-out-of-domain-robustness)
+- **READ ME PLEASE:** Here is a new paper from us describing challenges in robust pose estimation, why PRE-TRAINING really matters - which was our major scientific contribution to low-data input pose-estimation - and it describes new networks that are available to you. [Pretraining boosts out-of-domain robustness for pose estimation](https://paperswithcode.com/paper/pretraining-boosts-out-of-domain-robustness)
- - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/
+ - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/
- - **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
+- **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
+
- **REVIEW PAPER:** The state of animal pose estimation w/ deep learning i.e. "Deep learning tools for the measurement of animal behavior in neuroscience" [arXiv](https://arxiv.org/abs/1909.13868) & [published version](https://www.sciencedirect.com/science/article/pii/S0959438819301151)
- **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
-
- **WATCH:** There are a lot of docs... where to begin: [Video Tutorial!](https://www.youtube.com/watch?v=A9qZidI7tL8)
### **Module 1: getting started on data**
**What you need:** any videos where you can see the animals/objects, etc.
You can use our demo videos, grab some from the internet, or use whatever older data you have. Any camera, color/monochrome, etc will work. Find diverse videos, and label what you want to track well :)
-- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo 😊
- - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part1-labeling.pdf)
- - **READ ME PLEASE:** [DeepLabCut, the science](https://rdcu.be/4Rep)
- - **READ ME PLEASE:** [DeepLabCut, the user guide](https://rdcu.be/bHpHN)
- - **WATCH:** Video tutorial 1: [using the Project Manager GUI](https://www.youtube.com/watch?v=KcXogR-p5Ak)
- - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
- - **WATCH:** Video tutorial 2: [using the Project Manager GUI for multi-animal pose estimation](https://www.youtube.com/watch?v=Kp-stcTm77g)
- - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
- - **WATCH:** Video tutorial 3: [using ipython/pythonw (more functions!)](https://www.youtube.com/watch?v=7xwOhUcIGio)
- - multi-animal DLC: [labeling](https://www.youtube.com/watch?v=Kp-stcTm77g)
- - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
+- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo 😊
+ - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part1-labeling.pdf)
+ - **READ ME PLEASE:** [DeepLabCut, the science](https://rdcu.be/4Rep)
+ - **READ ME PLEASE:** [DeepLabCut, the user guide](https://rdcu.be/bHpHN)
+ - **WATCH:** Video tutorial 1: [using the Project Manager GUI](https://www.youtube.com/watch?v=KcXogR-p5Ak)
+ - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
+ - **WATCH:** Video tutorial 2: [using the Project Manager GUI for multi-animal pose estimation](https://www.youtube.com/watch?v=Kp-stcTm77g)
+ - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
+ - **WATCH:** Video tutorial 3: [using ipython/pythonw (more functions!)](https://www.youtube.com/watch?v=7xwOhUcIGio)
+ - multi-animal DLC: [labeling](https://www.youtube.com/watch?v=Kp-stcTm77g)
+ - Please go from project creation (use >1 video!) to labeling your data, and then check the labels!
### **Module 2: Neural Networks**
- - **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part2-network.pdf)
- - **READ ME PLEASE:** [What are convolutional neural networks?](https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
+- **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part2-network.pdf)
+
+- **READ ME PLEASE:** [What are convolutional neural networks?](https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
- - **READ ME PLEASE:** Here is a new paper from us describing challenges in robust pose estimation, why PRE-TRAINING really matters - which was our major scientific contribution to low-data input pose-estimation - and it describes new networks that are available to you. [Pretraining boosts out-of-domain robustness for pose estimation](https://paperswithcode.com/paper/pretraining-boosts-out-of-domain-robustness)
+- **READ ME PLEASE:** Here is a new paper from us describing challenges in robust pose estimation, why PRE-TRAINING really matters - which was our major scientific contribution to low-data input pose-estimation - and it describes new networks that are available to you. [Pretraining boosts out-of-domain robustness for pose estimation](https://paperswithcode.com/paper/pretraining-boosts-out-of-domain-robustness)
- - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/
+ - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/
- - **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
+- **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
+ -
-  +Before you create a training/test set, please read/watch:
- Before you create a training/test set, please read/watch:
- - **More information:** [Which types neural networks are available, and what should I use?](https://github.com/DeepLabCut/DeepLabCut/wiki/What-neural-network-should-I-use%3F-(Trade-offs,-speed-performance,-and-considerations))
- - **WATCH:** Video tutorial 1: [How to test different networks in a controlled way](https://www.youtube.com/watch?v=WXCVr6xAcCA)
- - Now, decide what model(s) you want to test.
- - IF you want to train on your CPU, then run the step `create_training_dataset`, in the GUI etc. on your own computer.
- - IF you want to use GPUs on google colab, [**(1)** watch this FIRST/follow along here!](https://www.youtube.com/watch?v=qJGs8nxx80A) **(2)** move your whole project folder to Google Drive, and then [**use this notebook**](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb)
+- **More information:** [Which types neural networks are available, and what should I use?](
+Before you create a training/test set, please read/watch:
- Before you create a training/test set, please read/watch:
- - **More information:** [Which types neural networks are available, and what should I use?](https://github.com/DeepLabCut/DeepLabCut/wiki/What-neural-network-should-I-use%3F-(Trade-offs,-speed-performance,-and-considerations))
- - **WATCH:** Video tutorial 1: [How to test different networks in a controlled way](https://www.youtube.com/watch?v=WXCVr6xAcCA)
- - Now, decide what model(s) you want to test.
- - IF you want to train on your CPU, then run the step `create_training_dataset`, in the GUI etc. on your own computer.
- - IF you want to use GPUs on google colab, [**(1)** watch this FIRST/follow along here!](https://www.youtube.com/watch?v=qJGs8nxx80A) **(2)** move your whole project folder to Google Drive, and then [**use this notebook**](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb)
+- **More information:** [Which types neural networks are available, and what should I use?](
- @@ -33,33 +30,44 @@ Now you have DeepLabCut installed, but if you want to update it, either follow t
Start at the Project Management Tab and work your way through the tabs to built your customized model and deploy it on new data.
We recommend to keep the terminal visible (as well as the GUI) so you can see the ongoing processes as you step through your project, or any errors that might arise.
-- For specific napari-based labeling features, see the ["napari gui" docs](napari-gui-usage).
+- For specific napari-based labeling features, see the {ref}`napari gui
@@ -33,33 +30,44 @@ Now you have DeepLabCut installed, but if you want to update it, either follow t
Start at the Project Management Tab and work your way through the tabs to built your customized model and deploy it on new data.
We recommend to keep the terminal visible (as well as the GUI) so you can see the ongoing processes as you step through your project, or any errors that might arise.
-- For specific napari-based labeling features, see the ["napari gui" docs](napari-gui-usage).
+- For specific napari-based labeling features, see the {ref}`napari gui
+Since visibility cannot be accounted for, it should be considered only a rough estimate of relative labeling progress rather than an absolute measure of completeness: hidden/occluded keypoints are not counted, therefore projects with occlusions will not have every body part on every individual in every frame.
+```
+
+### Point size slider
+
+The dock widget includes a slider to adjust the size of all keypoints in the viewer; the selected dot size will be saved in `config.yaml` for convenience, meaning DLC will reuse it for future sessions.
+
+## Copy-paste annotations
+
+To copy-paste keypoints from one frame to another:
+
+- Select the keypoints you want to copy using the selection tool (shortcut `3`)
+- Press `Ctrl+C` to copy the selected keypoints
+- Navigate to the target frame and press `Ctrl+V` to paste the keypoints
+
+## Color scheme display features
+
+The plugin shows a list of bodyparts and their corresponding colors in the dock widget. You can toggle the visibility of this color scheme using the **Show color scheme** button.
+
+```{tip}
+The display only shows keypoints that are currently visible in the viewer.
+To show all bodyparts in the color scheme from the config, use the checkbox at the top of the color scheme list.
+```
+
+### Quick body part/individual selection
+
+Clicking on a body part in the color scheme will select all keypoints of that body part in the viewer (including across individuals if applicable).
+
+This can be useful for quickly selecting and editing all keypoints of a specific body part.
+
+In individual coloring mode, the color scheme also shows the individuals list, and clicking on an individual will select all keypoints belonging to that individual.
+
+### Jump to body part in viewer
+
+To locate a bodypart label that is currently not visible in the viewer, enable "Show all bodyparts" in the color scheme list.
+Then, click on a bodypart entry in the color scheme list.
+The viewer will jump to the first instance of that body part and select it (when it exists).
+If the bodypart is already visible in the viewer, clicking on it in the color scheme will simply select all keypoints of that bodypart, as described above.
+
+This helps quickly find a specific body part in the viewer.
+
+## Trajectory plot
+
+The **Show trajectories** button opens a trajectory plot in a separate dock widget. This plot shows the trajectories of all **selected keypoints** over time, and will color-code them according to the active color scheme (bodyparts or individuals).
+
+To show the trajectory of a specific keypoint, simply select that keypoint in the viewer (using the selection tool or by clicking on the corresponding body part in the color scheme).
+
+Additional controls in the trajectory plot dock widget allow you to zoom and pan the plot, as well as adjust the time window shown.
diff --git a/docs/gui/napari/basic_usage.md b/docs/gui/napari/basic_usage.md
new file mode 100644
index 000000000..5e126b072
--- /dev/null
+++ b/docs/gui/napari/basic_usage.md
@@ -0,0 +1,285 @@
+---
+deeplabcut:
+ last_content_updated: '2026-04-09'
+ last_metadata_updated: '2026-04-09'
+ ignore: false
+ last_verified: '2026-04-09'
+ verified_for: 3.0.0rc14
+---
+
+(file:napari-dlc-basic-usage)=
+
+# napari-DLC - Basic usage
+
+`napari-deeplabcut` is a napari plugin for keypoint annotation and label refinement. It can be used either as part of the DeepLabCut GUI or as a standalone annotation tool.
+
+## Before you start
+
+If you installed `DeepLabCut[gui]`, `napari-deeplabcut` is already included.
+
+### In the DeepLabCut GUI
+
+When labeling frames, checking labels, or manually extracting frames from videos, the napari plugin will open automatically.
+
+### As a standalone plugin
+
+You can also install it as a standalone plugin:
+
+```bash
+pip install napari-deeplabcut
+```
+
+Start napari from a terminal:
+
+```bash
+napari
+```
+
+Then open the plugin from:
+
+**Plugins -> napari-deeplabcut: Keypoint controls**
+
+## Supported inputs
+
+The plugin reader can open the following inputs:
+
+- DeepLabCut `config.yaml`
+- Image folders (supports `.png`, `.jpg`, extracted frames from DLC, as well as folders of mixed formats)
+- Videos (`.mp4`, `.avi`, `.mov`)
+- `.h5` annotation files
+
+You can load files either by:
+
+- dragging and dropping them onto the napari viewer, or
+- using the **File** menu
+
+```{tip}
+If you drag and drop a compatible labeled-data folder, the widget opens automatically.
+```
+
+## Using napari
+
+```{important}
+To familiarize yourself with napari, we recommend checking out the [official napari documentation and tutorials](https://napari.org/stable/usage.html).
+```
+
+## Recommended basic labeling workflow
+
+The simplest way to **start labeling** is:
+
+1. Open an image-only folder
+1. Open the corresponding `config.yaml` from your DeepLabCut project
+
+**OR**
+
+1. Open a folder inside a DeepLabCut project's `labeled-data` directory with a `CollectedData_
+Keeping data inside the project directory is recommended for best compatibility. Fallbacks asking for the config file location are provided when this structure is not respected, but some features may be disabled or limited in that case.
+```
+
+### Useful shortcuts
+
+- napari native:
+ - `2` / `3`: switch between labeling and selection mode
+ - `4`: pan and zoom mode
+ - `Ctrl+R`: reset the viewer to the default zoom and position
+- napari-deeplabcut specific:
+ - `M`: cycle through annotation modes
+ - `E`: toggle edge coloring
+ - `F`: toggle between individual and bodypart coloring modes
+ - `V`: toggle visibility of the selected layer
+ - `Backspace`: delete selected point(s)
+ - `Ctrl+C` / `Ctrl+V`: copy and paste selected points
+
+```{tip}
+Use the **View shortcuts** button in the dock widget for a quick reference of napari-deeplabcut shortcuts and their context (i.e. when they are active).
+```
+
+### More quality-of-life features
+
+See the {ref}`Advanced features
+Make sure overwrite confirmation is enabled if you want to avoid accidentally overwriting existing `CollectedData...` annotations.
+```
+
+## Video workflow (crop and frame extraction)
+
+Videos can also be opened directly in napari.
+
+```{tip}
+This works best by using the main DLC GUI and following steps there for manual frame extraction, which will automatically open the video in napari.
+The workflow is otherwise the same when opening a video directly in napari.
+```
+
+When a video is loaded, the plugin provides a small video action panel that can be used to:
+
+- Extract the current frame into the dataset
+- Optionally export existing machine labels for that frame (load the corresponding h5 file first)
+- Define and save crop coordinates to the DeepLabCut `config.yaml`
+
+Keypoints from video-based workflows can be edited and saved in the same way as image-folder workflows.
+
+## Working with multiple folders
+
+We do not currently support working on **more than one dataset folder at a time**.
+If a new folder is opened while another one is already open, the plugin will prevent new frames from being loaded, attempt to load annotations using the current folder context, and show a warning.
+
+After finishing one folder, simply:
+
+1. Save the relevant **Points** layer
+1. Remove the current layers from the viewer using the layer list (left panel)
+1. Open the next folder (e.g. by dragging and dropping it onto the viewer)
+
+This helps keep saving behavior unambiguous.
+
+## Demo
+
+A short demo video is available here:
+
+[Link to video](https://youtu.be/hsA9IB5r73E)
+
+```{warning}
+This demo may be outdated, but the general annotation workflow remains the same. If you would like an updated video tutorial, please open a feature request issue on GitHub, and we will update it.
+
+```
diff --git a/docs/gui/napari_GUI.md b/docs/gui/napari_GUI.md
index 3c97d1f41..79f1ee7e9 100644
--- a/docs/gui/napari_GUI.md
+++ b/docs/gui/napari_GUI.md
@@ -1,228 +1,22 @@
---
deeplabcut:
last_content_updated: '2026-02-10'
- last_metadata_updated: '2026-03-06'
+ last_metadata_updated: '2026-04-09'
ignore: false
visibility: online
status: outdated
recommendation: archive
notes: Being updated in a separate PR (#3280)
+ last_verified: '2026-04-09'
+ verified_for: 3.0.0rc14
---
-(napari-gui)=
-# napari labeling GUI
+(file:napari-gui-landing)=
+# napari GUI
-We replaced wxPython with PySide6 + as of version 2.3. Here is how to use the napari-aspects of the new GUI. It is available in napari-hub as a stand alone GUI as well as integrated into our main GUI, [please see docs here](https://deeplabcut.github.io/DeepLabCut/docs/gui/PROJECT_GUI.html).
+Welcome to the documentation for napari-DLC, the napari plugin for keypoint annotation and label refinement. This plugin can be used either as part of the DeepLabCut GUI or as a standalone annotation tool.
-[](https://pypi.org/project/napari-deeplabcut)
-[](https://python.org)
-[](https://github.com/DeepLabCut/napari-deeplabcut/actions)
-[](https://codecov.io/gh/DeepLabCut/napari-deeplabcut)
-[](https://napari-hub.org/plugins/napari-deeplabcut)
+## Table of contents
-A napari plugin for keypoint annotation with DeepLabCut.
-
-
-## Installation
-
-You can install the full DeepLabCut napari-based GUI via [pip] by running this in your conda env:
-
-`pip install 'deeplabcut[tf,gui]'` or mac M1/M2 chip users: `pip install 'deeplabcut[apple_mchips,gui]'`
-
-*please note this is available since v2.3
-
-This is not needed if you ran the above installation, but you can install the stand-alone `napari-deeplabcut` via [pip]:
-
-` pip install napari-deeplabcut `
-
-
-To install latest development version:
-
- ` pip install git+https://github.com/DeepLabCut/napari-deeplabcut.git `
-
-
-(napari-gui-usage)=
-## Usage
-
-To use the full GUI, please run:
-
-`python -m deeplabcut`
-
-To use the stand-alone napari plugin, please launch napari:
-
-`napari `
-
-Then, activate the plugin in Plugins > napari-deeplabcut: Keypoint controls.
-
-All accepted files (`config.yaml`, images, `.h5` data files) can be loaded either by dropping them directly onto the canvas or via the File menu.
-
-The easiest way to get started is to drop a folder (typically a folder from within a DeepLabCut's `labeled-data` directory), and, if labeling from scratch, drop the corresponding `config.yaml` to automatically add a `Points layer` and populate the dropdown menus.
-
-[🎥 DEMO](https://youtu.be/hsA9IB5r73E)
-
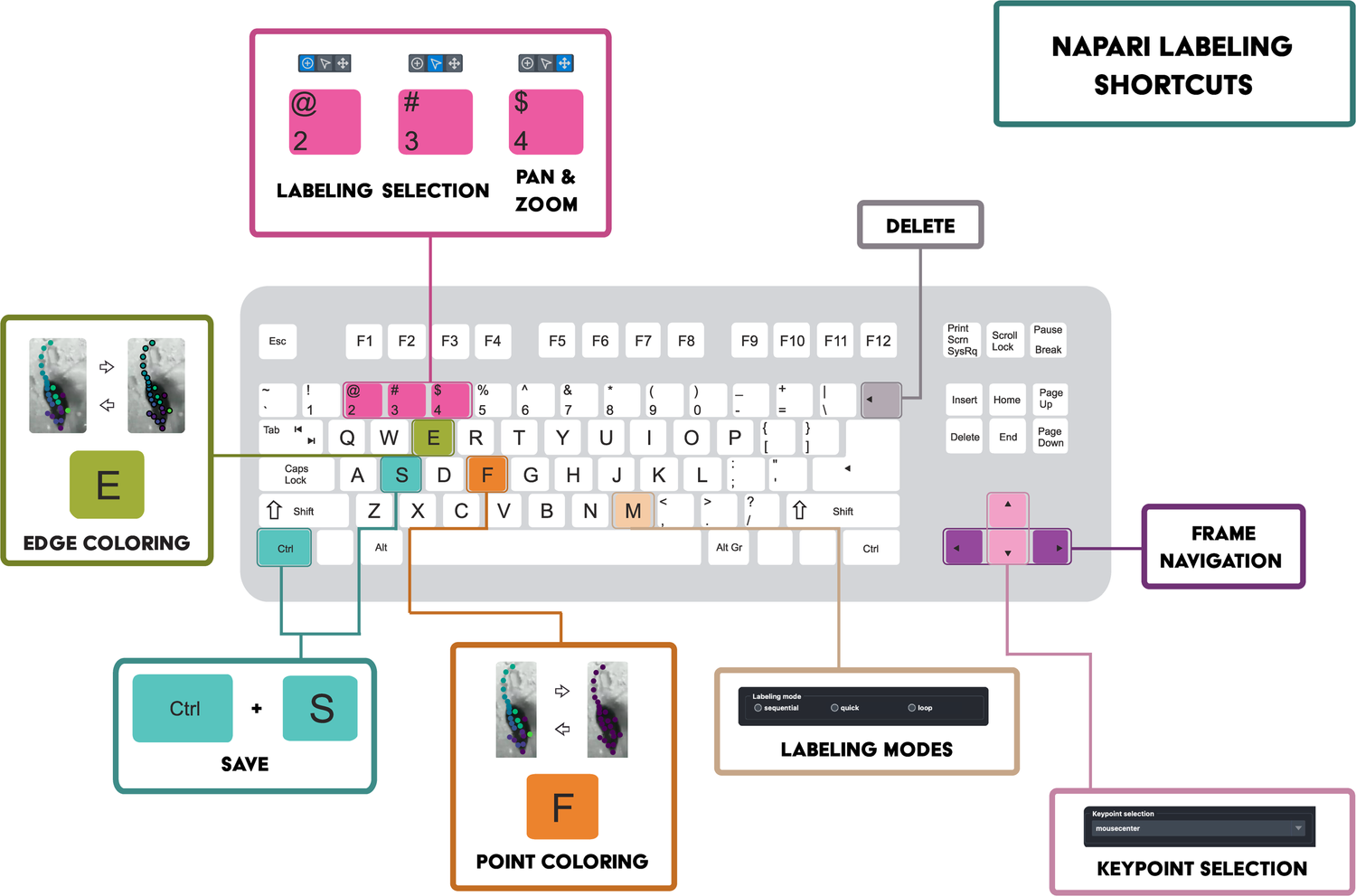
-**Tools & shortcuts are:**
-
-- `2` and `3`, to easily switch between labeling and selection mode
-- `4`, to enable pan & zoom (which is achieved using the mouse wheel or finger scrolling on the Trackpad)
-- `M`, to cycle through regular (sequential), quick, and cycle annotation mode (see the description [here](https://github.com/DeepLabCut/DeepLabCut-label/blob/ee71b0e15018228c98db3b88769e8a8f4e2c0454/dlclabel/layers.py#L9-L19))
-- `E`, to enable edge coloring (by default, if using this in refinement GUI mode, points with a confidence lower than 0.6 are marked
-in red)
-- `F`, to toggle between animal and body part color scheme.
-- `V`, to toggle visibility of the selected layer.
-- `backspace` to delete a point.
-- Check the box "display text" to show the label names on the canvas.
-- To move to another folder, be sure to save (Ctrl+S), then delete the layers, and re-drag/drop the next folder.
-
-
-
-
-
-### Save Layers
-
-Annotations and segmentations are saved with `File > Save Selected Layer(s)...` (or its shortcut `Ctrl+S`).
-Only when saving segmentation masks does a save file dialog pop up to name the destination folder;
-keypoint annotations are otherwise automatically saved in the corresponding folder as `CollectedData_ -
-  +**The installation process is as easy as the figure on the right!↘️**
-### 🚨 Before you start with our conda file, do you have a GPU?
-````{admonition} 🚨 Click here for more information!
-:class: dropdown
-- We recommend having a GPU if possible!
-- You **need to decide if you want to use a CPU or GPU for your models**: (Note, you can also use the CPU-only for project management and labeling the data! Then, for example, use Google Colaboratory GPUs for free (read more [here](https://github.com/DeepLabCut/DeepLabCut/tree/master/examples#demo-4-deeplabcut-training-and-analysis-on-google-colaboratory-with-googles-gpus) and there are a lot of helper videos on [our YouTube channel!](https://www.youtube.com/playlist?list=PLjpMSEOb9vRFwwgIkLLN1NmJxFprkO_zi)).
+### 🚨 Before you start...
- - **CPU?** Great, jump to the next section below!
+Do you have a GPU? If yes, see the {ref}`GPU support section
+**The installation process is as easy as the figure on the right!↘️**
-### 🚨 Before you start with our conda file, do you have a GPU?
-````{admonition} 🚨 Click here for more information!
-:class: dropdown
-- We recommend having a GPU if possible!
-- You **need to decide if you want to use a CPU or GPU for your models**: (Note, you can also use the CPU-only for project management and labeling the data! Then, for example, use Google Colaboratory GPUs for free (read more [here](https://github.com/DeepLabCut/DeepLabCut/tree/master/examples#demo-4-deeplabcut-training-and-analysis-on-google-colaboratory-with-googles-gpus) and there are a lot of helper videos on [our YouTube channel!](https://www.youtube.com/playlist?list=PLjpMSEOb9vRFwwgIkLLN1NmJxFprkO_zi)).
+### 🚨 Before you start...
- - **CPU?** Great, jump to the next section below!
+Do you have a GPU? If yes, see the {ref}`GPU support section
+
-- **Now, in Terminal (or Anaconda Command Prompt for Windows users)**, if you clicked to download, go to your downloads folder.
+- **Now, in Terminal (or Anaconda Command Prompt for Windows users)**:
-```{Hint}
-Windows users: Be sure to open the program terminal/cmd/anaconda prompt with a RIGHT-click, "open as admin"
-```
+ - If you clicked to download, go to your downloads folder.
-```{Hint}
-:class: dropdown
-If you cloned the repo onto your Desktop, the command may look like:
-``cd C:\Users\YourUserName\Desktop\DeepLabCut\conda-environments``
-You can (on Windows) hold SHIFT and right-click > Copy as path, or (on Mac) right-click and while in the menu press the OPTION key to reveal Copy as Pathname.
-```
-Be sure you are in the folder that has the `.yaml` file, then run:
+ - Be sure you are in the folder that has the `.yaml` file, then run:
+
+ `conda env create -f DEEPLABCUT.yaml`
+
+- You can now use this environment from anywhere on your computer.
+ Just activate your environment by running: `conda activate DEEPLABCUT`
-``conda env create -f DEEPLABCUT.yaml``
+Now you should see (`DEEPLABCUT`) on the left of your terminal screen:
+```
+(DEEPLABCUT) YourName-MacBook...
+```
-- You can now use this environment from anywhere on your computer (i.e., no need to go back into the conda- folder). Just enter your environment by running:
- - Ubuntu/MacOS: ``source/conda activate nameoftheenv`` (i.e. on your Mac: ``conda activate DEEPLABCUT``)
- - Windows: ``activate nameoftheenv`` (i.e. ``activate DEEPLABCUT``)
+```{note}
+No need to run `pip install deeplabcut`, it's already in the conda file!
+```
-Now you should see (`nameofenv`) on the left of your terminal screen, i.e. ``(DEEPLABCUT) YourName-MacBook...``
-NOTE: no need to run pip install deeplabcut, as it is already installed!!! :)
+(sec:deeplabcut-with-tf-install)=
-(deeplabcut-with-tf-install)=
-### 💡 Notice: PyTorch and TensorFlow Support within DeepLabCut
+#### TensorFlow support
````{admonition} DeepLabCut TensorFlow Support
-:class: dropdown
-As of June 2024 we have a PyTorch Engine backend and we will be depreciating the
-TensorFlow backend by the end of 2024. Currently, if you want to use TensorFlow, you
+---
+class: dropdown
+---
+💡 **PyTorch and TensorFlow Support within DeepLabCut**
+
+As of June 2024 we have a PyTorch Engine backend and we will be deprecating the
+TensorFlow backend by 2027.
+Currently, if you want to use TensorFlow, you
need to run `pip install deeplabcut[tf]` in order to install the correct version of
-TensorFlow in your conda env. Please note, we will be providing bug fixes, but we will
+TensorFlow in your conda env.
+Please note, we will be providing bug fixes, but we will
not be supporting new TensorFlow versions beyond 2.10 (Windows), and 2.12 for other OS.
Installing TensorFlow and getting it to have access to the GPU can be a bit tricky.
@@ -170,66 +200,78 @@ pip install --pre deeplabcut
```
````
-**Great, that's it! DeepLabCut is installed!** 🎉💜
+### Step 3: Let's run DeepLabCut!
+**DeepLabCut is installed!** 🎉💜
-### Step 3: Really, that's it! Let's run DeepLabCut
+Launch the DeepLabCut GUI in your new conda env by running `python -m deeplabcut`
Head over to the [User Guide Overview](https://deeplabcut.github.io/DeepLabCut/docs/UseOverviewGuide.html) for information.
-🎉 Launch DeepLabCut in your new env by running `python -m deeplabcut`
+```{warning}
+On **Windows**: Open the terminal/cmd/anaconda prompt as **Administrator** (right click and select "Run as administrator") to avoid permission issues when downloading models, and for symlink support when videos are not copied into the project folder.
+```
+
+### Conda environment management tips
+
+Here are some conda environment management tips: [kapeli.com: Conda Cheat Sheet](https://kapeli.com/cheat_sheets/Conda.docset/Contents/Resources/Documents/index)
+
+
-## Other ways to install DeepLabCut and additional tips
+
-### Alternatively, you can git clone this repo and install from source!
-i.e., if the download did not work or you just want to have the source code handy!
+Please see how to test your installation by following [this video](https://www.youtube.com/watch?v=IOWtKn3l33s).
+
+
+
+## Other ways to install DeepLabCut
+
+### git clone
+
+Recommended for users who want to modify the code, or want to be up-to-date with the latest code on GitHub.
- **Windows/Linux/MacBooks:** git clone this repo (in the terminal/cmd program, while **in a folder** you wish to place DeepLabCut
-To git clone type: ``git clone https://github.com/DeepLabCut/DeepLabCut.git``). Note, this can be anywhere, even downloads is fine.)
-- Then follow the same steps as in Step 2 above, adjusting for the file now being in the downloaded folder.
+- To git clone run: `git clone https://github.com/DeepLabCut/DeepLabCut.git`)
+- Then follow the same steps as in Step 2 above, adjusting for the `DEEPLABCUT.yaml` env file now being in the folder where you git cloned the repo.
+- Or use pip/uv to install from the cloned repo (see below).
-### PIP:
+(sec:uv-install)=
-- Everything you need to build custom models within DeepLabCut (i.e., use our source code and our dependencies) can be installed with `pip install 'deeplabcut[gui]'` (for GUI support w/PyTorch) or without the gui: `pip install 'deeplabcut'`.
-- If you want to use the SuperAnimal models, then please use `pip install 'deeplabcut[gui,modelzoo]'`.
+### `uv` (recommended for developers)
-## DOCKER:
+- Install `uv` following [instructions here](https://docs.astral.sh/uv/getting-started/installation/)
+- Run in the cloned repo:
-- We also have docker containers. Docker is the most reproducible way to use and deploy code. Please see our dedicated docker package and page [here](https://deeplabcut.github.io/DeepLabCut/docs/docker.html).
+```bash
+uv venv -p 3.12
+uv pip install -e .[gui,modelzoo,tf] # Change optional install as needed
+source .venv/bin/activate # or & .venv\Scripts\activate.ps1 on Windows
+```
-## Pro Tips:
+### `pip`
-More [installation ProTips](installation-tips) are also available.
+If you already have a local environment, everything you need to use the project manager GI, train and/or build custom models within DeepLabCut (i.e., use our source code and our dependencies) can be installed with `pip install 'deeplabcut[gui]'` (for GUI support w/PyTorch) or without the gui: `pip install 'deeplabcut'`.
-If you ever want to update your DLC, just run `pip install --upgrade deeplabcut` once
-you are inside your env. If you want to use a specific release, then you need to specify
-the version you want, such as `pip install deeplabcut==3.0`. Once installed, you can
-check the version by running `import deeplabcut` `deeplabcut.__version__`. Don't be
-afraid to update, DLC is backwards compatible with your 2.0+ projects and performance
-continues to get better and new features are added nearly monthly.
+- If you **cloned the repo** and want to make edits to the code locally, navigate to the cloned repo folder and run `pip install -e .[gui,modelzoo,tf]` to install the package in "editable" mode, which allows you to make changes to the code and have those changes reflected when you import the package.
+- If you want to use the SuperAnimal models, then please use `pip install 'deeplabcut[gui,modelzoo]'`.
-**All of the data you labelled in version 2.X is also compatible with version 3+ and the
-PyTorch engine**! There is no change in the workflow or the way labels are handled: the
-big changes happen under-the-hood! If you've been working with DeepLabCut 2.X and want
-to learn more about moving to the PyTorch engine, checkout our docs on [moving from
-TensorFlow to PyTorch](dlc3-user-guide)
+### Docker
-Here are some conda environment management tips: [kapeli.com: Conda Cheat Sheet](
-https://kapeli.com/cheat_sheets/Conda.docset/Contents/Resources/Documents/index)
+- We also have docker containers. Docker is the most reproducible way to use and deploy code. Please see our dedicated docker package and page [here](https://deeplabcut.github.io/DeepLabCut/docs/docker.html).
-**Pro Tip:** If you want to modify code and then test it, you can use our provided
-testscripts. This would mean you need to be up-to-date with the latest GitHub-based code
-though! Please see [here](installation-tips) on how to get the latest GitHub code, and
-how to test your installation by following this video:
-https://www.youtube.com/watch?v=IOWtKn3l33s.
+### Creating your own conda environment
-## Creating your own customized conda env (recommended route for Linux: Ubuntu, CentOS, Mint, etc.)
+
-*Note in a fresh ubuntu install, you will often have to run: ``sudo apt-get install gcc python3-dev`` to install the GNU Compiler Collection and the python developing environment.
+
-Some users might want to create their own customize env. - Here is an example.
+```{tip}
+In a fresh ubuntu install, you will often have to run: `sudo apt-get install gcc python3-dev` to install the GNU Compiler Collection and the python developing environment.
+```
-In the terminal type:
+Create a new conda environment with Python 3.10 (or 3.11, 3.12) by running:
`conda create -n DLC python=3.10`
@@ -237,68 +279,87 @@ In the terminal type:
`pip install deeplabcut`) or `pip install 'deeplabcut[gui]'` which has a napari based
GUI.
+## Updating your installation
+
+If you ever want to update your DLC, just run `pip install --upgrade deeplabcut` inside your env.
+If you want to use a specific release, then specify the version you want, such as `pip install deeplabcut==3.0`.
+Once installed, you can
+check the version by running `import deeplabcut` `deeplabcut.__version__`.
+
+Don't be afraid to update, DLC is backwards compatible with your 2.0+ projects and performance continues to get better and new features are added often.
+
+### Data compatibility
+
+**All of the data you labelled in version 2.X is also compatible with version 3+ and the
+PyTorch engine**!
+There is no change in the workflow or the way labels are handled: the
+big changes happen under-the-hood! If you've been working with DeepLabCut 2.X and want
+to learn more about moving to the PyTorch engine, check out our docs on [moving from
+TensorFlow to PyTorch](dlc3-user-guide)
+
+(sec:install-gpu-support)=
-## **GPU Support:**
+## GPU Support
-The ONLY thing you need to do **first** if you have an NVIDIA GPU and the matching NVIDIA CUDA+driver installed.
-- CUDA: https://developer.nvidia.com/cuda-downloads (just follow the prompts here!)
-- DRIVERS: https://www.nvidia.com/Download/index.aspx
+### General GPU support
-### The most common "new user" hurdle is installing and using your GPU, so don't get discouraged!
+Please ensure you have an NVIDIA GPU and the matching NVIDIA driver installed.
-**CRITICAL:** If you have a GPU, you should FIRST **install an appropriate driver for
-your specific GPU**, then you can use the supplied conda file. You'll need an NVIDIA GPU
-which is compatible with CUDA. To see a list of CUDA-enabled NVIDIA GPUs, please [see
-their website](https://developer.nvidia.com/cuda-gpus).
+```{warning}
+If you have a GPU, you should first **install an appropriate driver for
+your specific GPU**, then you can use the supplied conda file.
+```
-- Here we provide notes on how to install and check your GPU use with TensorFlow (which
-is used by DeepLabCut and already installed with the Anaconda files above). Thus, you do
-not need to independently install tensorflow.
+- Drivers: see [NVIDIA Drivers](https://www.nvidia.com/Download/index.aspx)
+- CUDA: download [here](https://developer.nvidia.com/cuda-downloads) if needed. Installing the drivers usually allows you to skip installing CUDA; instead obtaining via the PyTorch installation process.
-**FIRST**, install a driver for your GPU. Find DRIVER HERE:
-https://www.nvidia.com/download/index.aspx
+### Installing CUDA and cuDNN for TensorFlow GPU support
-- Check which driver is installed by typing this into the terminal: ``nvidia-smi``.
+You will need an NVIDIA GPU that is compatible with CUDA.
-**SECOND**, install CUDA: https://developer.nvidia.com/ (Note that cuDNN, https://developer.nvidia.com/cudnn, is supplied inside the anaconda environment files, so you don't need to install it again).
+To see a list of CUDA-enabled NVIDIA GPUs, please [see their website](https://developer.nvidia.com/cuda-gpus).
-**THIRD:** Follow the steps above to get the `DEEPLABCUT` conda file and install it!
+Here we provide notes on how to install and check your GPU use with TensorFlow, which is used by DeepLabCut and will be installed with the Anaconda files above.
+Thus, you do not need to independently install tensorflow.
-### Notes:
+1. Install a driver for your GPU, using the NVIDIA Drivers link above.
+ - Check which driver is installed by typing this into the terminal: `nvidia-smi`.
+1. Install [CUDA](https://developer.nvidia.com/). Note that [cuDNN](https://developer.nvidia.com/cudnn) is supplied inside the anaconda environment files, so you don't need to install it again.
+1. Follow the steps above to get the `DEEPLABCUT` conda file and install it!
+
+### Notes
- **As of version 3.0+ we moved to PyTorch. The Last supported version of TensorFlow is
-2.10 (window users) and 2.12 for others (we have not tested beyond this).**
+ 2.10 (window users) and 2.12 for others** (We will not be testing beyond this)
+
- Please be mindful different versions of TensorFlow require different CUDA versions.
+
- As the combination of TensorFlow and CUDA matters, we strongly encourage you to
-**check your driver/cuDNN/CUDA/TensorFlow versions** [on this StackOverflow post](
-https://stackoverflow.com/questions/30820513/what-is-version-of-cuda-for-nvidia-304-125/30820690#30820690
-).
+ **check your driver/cuDNN/CUDA/TensorFlow versions** [on this StackOverflow post](https://stackoverflow.com/questions/30820513/what-is-version-of-cuda-for-nvidia-304-125/30820690#30820690).
+
- To check your GPU is working, in the terminal, run:
-`nvcc -V` to check your installed version(s).
+ `nvcc -V` to check your installed version(s).
- The best practice is to then run the supplied `testscript_pytorch_single_animal.py`
-(or `testscript_tensorflow_single_animal.py` for the TensorFlow engine); this is inside the examples folder you
-acquired when you git cloned the repo. Here is more information/a short
-[video on running the testscript](https://www.youtube.com/watch?v=IOWtKn3l33s).
-- Additionally, if you want to use the bleeding edge, with your git clone you also get
-the latest code. While inside the main DeepLabCut folder, you can run `./reinstall.sh`
-to be sure it's installed (more [here](installation-tips))
-- You can test that your GPU is being properly engaged with these additional [tips](
-https://www.tensorflow.org/programmers_guide/using_gpu).
-- Ubuntu users might find this [installation guide](
-https://deeplabcut.github.io/DeepLabCut/docs/recipes/installTips.html#installation-on-ubuntu-20-04-lts
-) for a fresh ubuntu install useful as well.
-
-## Troubleshooting:
-
-TensorFlow:
+ (or `testscript_tensorflow_single_animal.py` for the TensorFlow engine); this is inside the examples folder you
+ acquired when you git cloned the repo. Here is more information/a short
+ [video on running the test scripts](https://www.youtube.com/watch?v=IOWtKn3l33s).
+
+- You can test that your GPU is being properly used with these additional [tips](https://www.tensorflow.org/programmers_guide/using_gpu).
+
+- Ubuntu users might find this [installation guide](https://deeplabcut.github.io/DeepLabCut/docs/recipes/installTips.html#installation-on-ubuntu-20-04-lts) for a fresh DLC install on Ubuntu useful as well.
+
+## Troubleshooting
+
+### TensorFlow
+
Here are some additional resources users have found helpful (posted without endorsement):
- https://stackoverflow.com/questions/30820513/what-is-the-correct-version-of-cuda-for-my-nvidia-driver/30820690
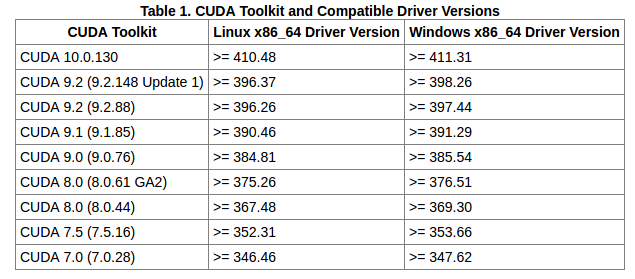 +
+ 