
 ## **ATTENTION: Our code base in this repo assumes you:**
-A. You have 2D videos and a DeepLabCut network to analyze them as described in the [main documentation](overview). This can be with multiple separate networks for each camera (less recommended), or one network trained on all views - recommended! (See [Nath*, Mathis* et al., 2019](https://www.biorxiv.org/content/10.1101/476531v1)). We also support multi-animal 3D with this code (please see [Lauer et al. 2022](https://doi.org/10.1038/s41592-022-01443-0)).
+A. You have 2D videos and a DeepLabCut network to analyze them as described in the
+[main documentation](overview). This can be with multiple
+separate networks for each camera (less recommended), or one network trained on all views - recommended! (See
+[Nath*, Mathis* et al., 2019](https://www.biorxiv.org/content/10.1101/476531v1)). We also support multi-animal 3D with this code (please see
+[Lauer et al. 2022](https://doi.org/10.1038/s41592-022-01443-0)).
B. You are using 2 cameras, in a [stereo configuration](https://github.com/DeepLabCut/DeepLabCut/blob/5ac4c8cb6bcf2314a3abfcf979b8dd170608e094/deeplabcut/pose_estimation_3d/camera_calibration.py#L223), for 3D*.
@@ -20,12 +27,16 @@ Here are other excellent options for you to use that extend DeepLabCut:
## **ATTENTION: Our code base in this repo assumes you:**
-A. You have 2D videos and a DeepLabCut network to analyze them as described in the [main documentation](overview). This can be with multiple separate networks for each camera (less recommended), or one network trained on all views - recommended! (See [Nath*, Mathis* et al., 2019](https://www.biorxiv.org/content/10.1101/476531v1)). We also support multi-animal 3D with this code (please see [Lauer et al. 2022](https://doi.org/10.1038/s41592-022-01443-0)).
+A. You have 2D videos and a DeepLabCut network to analyze them as described in the
+[main documentation](overview). This can be with multiple
+separate networks for each camera (less recommended), or one network trained on all views - recommended! (See
+[Nath*, Mathis* et al., 2019](https://www.biorxiv.org/content/10.1101/476531v1)). We also support multi-animal 3D with this code (please see
+[Lauer et al. 2022](https://doi.org/10.1038/s41592-022-01443-0)).
B. You are using 2 cameras, in a [stereo configuration](https://github.com/DeepLabCut/DeepLabCut/blob/5ac4c8cb6bcf2314a3abfcf979b8dd170608e094/deeplabcut/pose_estimation_3d/camera_calibration.py#L223), for 3D*.
@@ -20,12 +27,16 @@ Here are other excellent options for you to use that extend DeepLabCut:
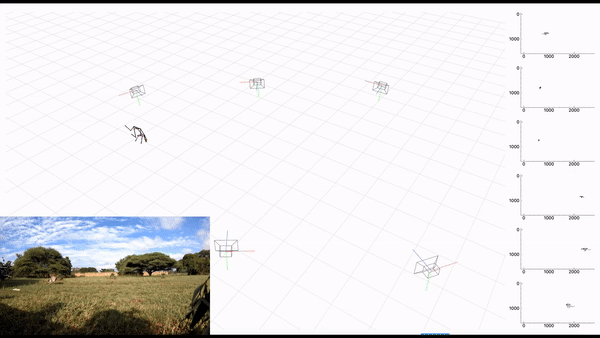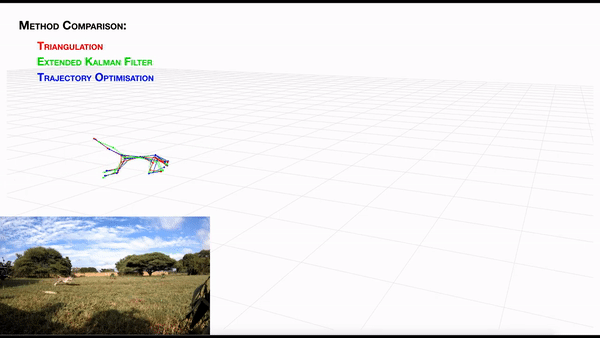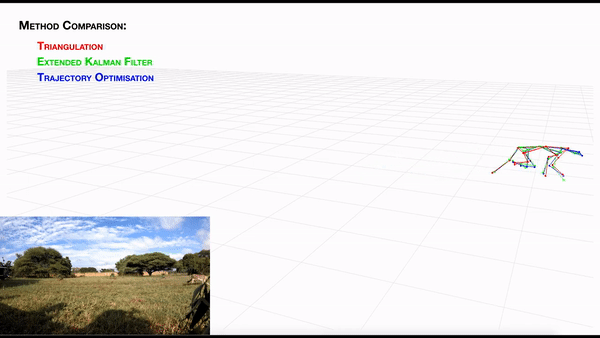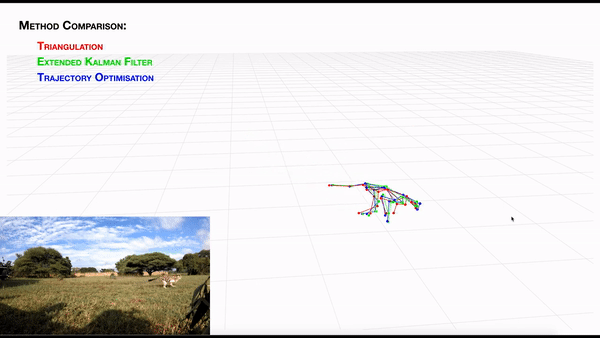 -- **[AcinoSet](https://github.com/African-Robotics-Unit/AcinoSet)**; **n**-camera support with triangulation, extended Kalman filtering, and trajectory optimization code (see video to the right for a min demo, courtesy of Prof. Patel), plus a GUI to visualize 3D data. It is built to work directly with DeepLabCut (but currently tailored to cheetah's, thus some coding skills are required at this time).
+- **[AcinoSet](https://github.com/African-Robotics-Unit/AcinoSet)**; **n**-camera support with triangulation, extended Kalman filtering, and trajectory optimization
+code (see video to the right for a min demo, courtesy of Prof. Patel), plus a GUI to visualize 3D data. It is built to
+work directly with DeepLabCut (but currently tailored to cheetah's, thus some coding skills are required at this time).
-- **[anipose.org](https://anipose.readthedocs.io/en/latest/)**; a wrapper for 3D deeplabcut that provides >3 camera support and is built to work directly with DeepLabCut. You can `pip install anipose` into your DLC conda environment.
+- **[anipose.org](https://anipose.readthedocs.io/en/latest/)**; a wrapper for 3D deeplabcut that provides >3 camera support and is built to work directly with
+DeepLabCut. You can `pip install anipose` into your DLC conda environment.
-- **Argus, easywand or DLTdv** w/DeepLabCut see https://github.com/haliaetus13/DLCconverterDLT; this can be used with the the highly popular Argus or DLTdv tools for wand calibration.
+- **Argus, easywand or DLTdv** w/DeepLabCut see https://github.com/haliaetus13/DLCconverterDLT; this can be used with
+the the highly popular Argus or DLTdv tools for wand calibration.
## Jump in with direct DeepLabCut 2-camera support:
@@ -39,83 +50,118 @@ Here are other excellent options for you to use that extend DeepLabCut:
### (1) Create a New 3D Project:
-Watch a [DEMO VIDEO](https://youtu.be/Eh6oIGE4dwI) on how to use this code, and check out the Notebook [here](https://github.com/DeepLabCut/DeepLabCut/blob/master/examples/JUPYTER/Demo_3D_DeepLabCut.ipynb)!
+Watch a [DEMO VIDEO](https://youtu.be/Eh6oIGE4dwI) on how to use this code, and check out the Notebook [here](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/JUPYTER/Demo_3D_DeepLabCut.ipynb)!
-You will run this function **one** time per project; a project is defined as a given set of cameras and calibration images. You can always analyze new videos within this project.
+You will run this function **one** time per project; a project is defined as a given set of cameras and calibration
+images. You can always analyze new videos within this project.
-The function **create\_new\_project\_3d** creates a new project directory specifically for converting the 2D pose to 3D pose, required subdirectories, and a basic 3D project configuration file. Each project is identified by the name of the project (e.g. Task1), name of the experimenter (e.g. YourName), as well as the date at creation.
+The function **create\_new\_project\_3d** creates a new project directory specifically for converting the 2D pose to 3D
+pose, required subdirectories, and a basic 3D project configuration file. Each project is identified by the name of the
+project (e.g. Task1), name of the experimenter (e.g. YourName), as well as the date at creation.
-Thus, this function requires the user to input the enter the name of the project, the name of the experimenter and number of cameras to be used. Currently, DeepLabCut supports triangulation using 2 cameras, but will expand to more than 2 cameras in a future version.
+Thus, this function requires the user to enter the name of the project, the name of the experimenter and number of
+cameras to be used. Currently, DeepLabCut supports triangulation using 2 cameras, but will expand to more than 2 cameras
+in a future version.
To start a 3D project type the following in ipython:
```python
-deeplabcut.create_new_project_3d('ProjectName','NameofLabeler',num_cameras = 2)
+deeplabcut.create_new_project_3d("ProjectName", "NameofLabeler", num_cameras=2)
```
-TIP 1: you can also pass ``working_directory=`Full path of the working directory'`` if you want to place this folder somewhere beside the current directory you are working in. If the optional argument ``working_directory`` is unspecified, the project directory is created in the current working directory.
+TIP 1: you can also pass `working_directory="Full path of the working directory"` if you want to place this folder
+somewhere beside the current directory you are working in. If the optional argument `working_directory` is unspecified,
+the project directory is created in the current working directory.
-TIP 2: you can also place ``config_path3d`` in front of ``deeplabcut.create_new_project_3d`` to create a variable that holds the path to the config.yaml file, i.e. ``config_path3d=deeplabcut.create_new_project_3d(...`` Or, set this variable for easy use. Please note that ``config_path3d='Full path of the 3D project configuration file'``.
+TIP 2: you can also place `config_path3d` in front of `deeplabcut.create_new_project_3d` to create a variable that holds
+the path to the config.yaml file, i.e. `config_path3d=deeplabcut.create_new_project_3d(...` Or, set this variable for
+easy use. Please note that `config_path3d='Full path of the 3D project configuration file'`.
- This function will create a project directory with the name **Name of the project+name of the experimenter+date of creation of the project+3d** in the **Working directory**. The project directory will have subdirectories: **calibration_images**, **camera_matrix**, **corners**, and **undistortion**. All the outputs generated during the course of a project will be stored in one of these subdirectories, thus allowing each project to be curated in separation from other projects.
+This function will create a project directory with the name **Name of the project+name of the experimenter+date of
+creation of the project+3d** in the **Working directory**. The project directory will have subdirectories:
+**calibration_images**, **camera_matrix**, **corners**, and **undistortion**. All the outputs generated during the
+course of a project will be stored in one of these subdirectories, thus allowing each project to be curated in
+separation from other projects.
- The purpose of the subdirectories is as follows:
+The purpose of the subdirectories is as follows:
- **calibration_images:** This directory will contain a set of calibration images acquired from the two cameras. A calibration image can be acquired using a printed checkerboard and its pair wise images are taken from both the cameras to consider as a set of calibration images. These pair of images are saved as ``.jpg`` with camera names as the prefix. e.g. ``camera-1-01.jpg`` and ``camera-2-01.jpg`` for the first pair of images. While taking the images:
-- Keep the orientation of the chessboard same and do not rotate more than 30 degrees. Rotating the chessboard circular will change the origin across the frames and may result in incorrect order of detected corners.
-- Cover several distances, and within each distance, cover all parts of the image view (all corners and center).
-Use a chessboard as big as possible, ideally a chessboard with of at least 8x6 squares.
-- Aim for taking at least 70 pair of images as after corner detection, some of the images might need to be discarded due to either incorrect corner detection or incorrect order of detected corners.
+**calibration_images:** This directory will contain a set of calibration images acquired from the two cameras. A
+calibration image can be acquired using a printed checkerboard and its pair wise images are taken from both the cameras
+to consider as a set of calibration images.
- **camera_matrix:** This directory will store the parameter for both the cameras as a pickle file. Specifically, these pickle files contain the intrinsic and extrinsic camera parameters. While the intrinsic parameters represent a transformation from 3-D camera's coordinates into the image coordinates, the extrinsic parameters represent a rigid transformation from world coordinate system to the 3-D camera's coordinate system.
+**camera_matrix:** This directory will store the parameter for both the cameras as a pickle file. Specifically, these
+pickle files contain the intrinsic and extrinsic camera parameters. While the intrinsic parameters represent a
+transformation from 3-D camera's coordinates into the image coordinates, the extrinsic parameters represent a rigid
+transformation from world coordinate system to the 3-D camera's coordinate system.
- **corners:** As a part of camera calibration, the checkerboard pattern is detected in the calibration images and these patterns will be stored in this directory. Each row of the checkerboard grid is marked with a unique color.
+**corners:** As a part of camera calibration, the checkerboard pattern is detected in the calibration images and these
+patterns will be stored in this directory. Each row of the checkerboard grid is marked with a unique color.
- **undistortion:** In order to check for calibration, the calibration images and the corresponding corner points are undistorted. These undistorted images are overlaid with undistorted points and will be stored in this directory.
+**undistortion:** In order to check for calibration, the calibration images and the corresponding corner points are
+undistorted. These undistorted images are overlaid with undistorted points and will be stored in this directory.
- Here is an overview of the calibration and triangulation workflow that follows:
+Here is an overview of the calibration and triangulation workflow that follows:
-
-- **[AcinoSet](https://github.com/African-Robotics-Unit/AcinoSet)**; **n**-camera support with triangulation, extended Kalman filtering, and trajectory optimization code (see video to the right for a min demo, courtesy of Prof. Patel), plus a GUI to visualize 3D data. It is built to work directly with DeepLabCut (but currently tailored to cheetah's, thus some coding skills are required at this time).
+- **[AcinoSet](https://github.com/African-Robotics-Unit/AcinoSet)**; **n**-camera support with triangulation, extended Kalman filtering, and trajectory optimization
+code (see video to the right for a min demo, courtesy of Prof. Patel), plus a GUI to visualize 3D data. It is built to
+work directly with DeepLabCut (but currently tailored to cheetah's, thus some coding skills are required at this time).
-- **[anipose.org](https://anipose.readthedocs.io/en/latest/)**; a wrapper for 3D deeplabcut that provides >3 camera support and is built to work directly with DeepLabCut. You can `pip install anipose` into your DLC conda environment.
+- **[anipose.org](https://anipose.readthedocs.io/en/latest/)**; a wrapper for 3D deeplabcut that provides >3 camera support and is built to work directly with
+DeepLabCut. You can `pip install anipose` into your DLC conda environment.
-- **Argus, easywand or DLTdv** w/DeepLabCut see https://github.com/haliaetus13/DLCconverterDLT; this can be used with the the highly popular Argus or DLTdv tools for wand calibration.
+- **Argus, easywand or DLTdv** w/DeepLabCut see https://github.com/haliaetus13/DLCconverterDLT; this can be used with
+the the highly popular Argus or DLTdv tools for wand calibration.
## Jump in with direct DeepLabCut 2-camera support:
@@ -39,83 +50,118 @@ Here are other excellent options for you to use that extend DeepLabCut:
### (1) Create a New 3D Project:
-Watch a [DEMO VIDEO](https://youtu.be/Eh6oIGE4dwI) on how to use this code, and check out the Notebook [here](https://github.com/DeepLabCut/DeepLabCut/blob/master/examples/JUPYTER/Demo_3D_DeepLabCut.ipynb)!
+Watch a [DEMO VIDEO](https://youtu.be/Eh6oIGE4dwI) on how to use this code, and check out the Notebook [here](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/JUPYTER/Demo_3D_DeepLabCut.ipynb)!
-You will run this function **one** time per project; a project is defined as a given set of cameras and calibration images. You can always analyze new videos within this project.
+You will run this function **one** time per project; a project is defined as a given set of cameras and calibration
+images. You can always analyze new videos within this project.
-The function **create\_new\_project\_3d** creates a new project directory specifically for converting the 2D pose to 3D pose, required subdirectories, and a basic 3D project configuration file. Each project is identified by the name of the project (e.g. Task1), name of the experimenter (e.g. YourName), as well as the date at creation.
+The function **create\_new\_project\_3d** creates a new project directory specifically for converting the 2D pose to 3D
+pose, required subdirectories, and a basic 3D project configuration file. Each project is identified by the name of the
+project (e.g. Task1), name of the experimenter (e.g. YourName), as well as the date at creation.
-Thus, this function requires the user to input the enter the name of the project, the name of the experimenter and number of cameras to be used. Currently, DeepLabCut supports triangulation using 2 cameras, but will expand to more than 2 cameras in a future version.
+Thus, this function requires the user to enter the name of the project, the name of the experimenter and number of
+cameras to be used. Currently, DeepLabCut supports triangulation using 2 cameras, but will expand to more than 2 cameras
+in a future version.
To start a 3D project type the following in ipython:
```python
-deeplabcut.create_new_project_3d('ProjectName','NameofLabeler',num_cameras = 2)
+deeplabcut.create_new_project_3d("ProjectName", "NameofLabeler", num_cameras=2)
```
-TIP 1: you can also pass ``working_directory=`Full path of the working directory'`` if you want to place this folder somewhere beside the current directory you are working in. If the optional argument ``working_directory`` is unspecified, the project directory is created in the current working directory.
+TIP 1: you can also pass `working_directory="Full path of the working directory"` if you want to place this folder
+somewhere beside the current directory you are working in. If the optional argument `working_directory` is unspecified,
+the project directory is created in the current working directory.
-TIP 2: you can also place ``config_path3d`` in front of ``deeplabcut.create_new_project_3d`` to create a variable that holds the path to the config.yaml file, i.e. ``config_path3d=deeplabcut.create_new_project_3d(...`` Or, set this variable for easy use. Please note that ``config_path3d='Full path of the 3D project configuration file'``.
+TIP 2: you can also place `config_path3d` in front of `deeplabcut.create_new_project_3d` to create a variable that holds
+the path to the config.yaml file, i.e. `config_path3d=deeplabcut.create_new_project_3d(...` Or, set this variable for
+easy use. Please note that `config_path3d='Full path of the 3D project configuration file'`.
- This function will create a project directory with the name **Name of the project+name of the experimenter+date of creation of the project+3d** in the **Working directory**. The project directory will have subdirectories: **calibration_images**, **camera_matrix**, **corners**, and **undistortion**. All the outputs generated during the course of a project will be stored in one of these subdirectories, thus allowing each project to be curated in separation from other projects.
+This function will create a project directory with the name **Name of the project+name of the experimenter+date of
+creation of the project+3d** in the **Working directory**. The project directory will have subdirectories:
+**calibration_images**, **camera_matrix**, **corners**, and **undistortion**. All the outputs generated during the
+course of a project will be stored in one of these subdirectories, thus allowing each project to be curated in
+separation from other projects.
- The purpose of the subdirectories is as follows:
+The purpose of the subdirectories is as follows:
- **calibration_images:** This directory will contain a set of calibration images acquired from the two cameras. A calibration image can be acquired using a printed checkerboard and its pair wise images are taken from both the cameras to consider as a set of calibration images. These pair of images are saved as ``.jpg`` with camera names as the prefix. e.g. ``camera-1-01.jpg`` and ``camera-2-01.jpg`` for the first pair of images. While taking the images:
-- Keep the orientation of the chessboard same and do not rotate more than 30 degrees. Rotating the chessboard circular will change the origin across the frames and may result in incorrect order of detected corners.
-- Cover several distances, and within each distance, cover all parts of the image view (all corners and center).
-Use a chessboard as big as possible, ideally a chessboard with of at least 8x6 squares.
-- Aim for taking at least 70 pair of images as after corner detection, some of the images might need to be discarded due to either incorrect corner detection or incorrect order of detected corners.
+**calibration_images:** This directory will contain a set of calibration images acquired from the two cameras. A
+calibration image can be acquired using a printed checkerboard and its pair wise images are taken from both the cameras
+to consider as a set of calibration images.
- **camera_matrix:** This directory will store the parameter for both the cameras as a pickle file. Specifically, these pickle files contain the intrinsic and extrinsic camera parameters. While the intrinsic parameters represent a transformation from 3-D camera's coordinates into the image coordinates, the extrinsic parameters represent a rigid transformation from world coordinate system to the 3-D camera's coordinate system.
+**camera_matrix:** This directory will store the parameter for both the cameras as a pickle file. Specifically, these
+pickle files contain the intrinsic and extrinsic camera parameters. While the intrinsic parameters represent a
+transformation from 3-D camera's coordinates into the image coordinates, the extrinsic parameters represent a rigid
+transformation from world coordinate system to the 3-D camera's coordinate system.
- **corners:** As a part of camera calibration, the checkerboard pattern is detected in the calibration images and these patterns will be stored in this directory. Each row of the checkerboard grid is marked with a unique color.
+**corners:** As a part of camera calibration, the checkerboard pattern is detected in the calibration images and these
+patterns will be stored in this directory. Each row of the checkerboard grid is marked with a unique color.
- **undistortion:** In order to check for calibration, the calibration images and the corresponding corner points are undistorted. These undistorted images are overlaid with undistorted points and will be stored in this directory.
+**undistortion:** In order to check for calibration, the calibration images and the corresponding corner points are
+undistorted. These undistorted images are overlaid with undistorted points and will be stored in this directory.
- Here is an overview of the calibration and triangulation workflow that follows:
+Here is an overview of the calibration and triangulation workflow that follows:
- +


 @@ -213,22 +296,36 @@ The **triangulated file** is now saved under the same directory where the video
### (5) Visualize your 3D DeepLabCut Videos:
-In order to visualize both the 2D videos with tracked points plut the pose in 3D, the user can create a 3D video for certain frames (these are large files, so we advise just looking at a subset of frames). The user can specify the config file, the **path of the triangulated file folder**, and specify the start and end frame indices to create a 3D labeled video. Note that the ``triangulated_file_folder`` is where the newly created file that ends with ``yourDLC_3D_scorername.h5`` is located. This can be done using:
+In order to visualize both the 2D videos with tracked points plut the pose in 3D, the user can create a 3D video for
+certain frames (these are large files, so we advise just looking at a subset of frames). The user can specify the config
+file, the **path of the triangulated file folder**, and specify the start and end frame indices to create a 3D labeled
+video. Note that the `triangulated_file_folder` is where the newly created file that ends with
+`yourDLC_3D_scorername.h5` is located. This can be done using:
```python
-deeplabcut.create_labeled_video_3d(config_path, ['triangulated_file_folder'], start=50, end=250)
+deeplabcut.create_labeled_video_3d(
+ config_path,
+ ["triangulated_file_folder"],
+ start=50,
+ end=250
+)
```
-**TIP:** (see more parameters below) You can set how the axis of the 3D plot on the far right looks by changing the variables ``xlim``, ``ylim``, ``zlim`` and ``view``. Your checkerboard_3d.png image which was created above will show you the axis ranges. Here is an example:
+**TIP:** (see more parameters below) You can set how the axis of the 3D plot on the far right looks by changing the
+variables `xlim`, `ylim`, `zlim` and `view`. Your checkerboard_3d.png image which was created above will show you the
+axis ranges. Here is an example:
@@ -213,22 +296,36 @@ The **triangulated file** is now saved under the same directory where the video
### (5) Visualize your 3D DeepLabCut Videos:
-In order to visualize both the 2D videos with tracked points plut the pose in 3D, the user can create a 3D video for certain frames (these are large files, so we advise just looking at a subset of frames). The user can specify the config file, the **path of the triangulated file folder**, and specify the start and end frame indices to create a 3D labeled video. Note that the ``triangulated_file_folder`` is where the newly created file that ends with ``yourDLC_3D_scorername.h5`` is located. This can be done using:
+In order to visualize both the 2D videos with tracked points plut the pose in 3D, the user can create a 3D video for
+certain frames (these are large files, so we advise just looking at a subset of frames). The user can specify the config
+file, the **path of the triangulated file folder**, and specify the start and end frame indices to create a 3D labeled
+video. Note that the `triangulated_file_folder` is where the newly created file that ends with
+`yourDLC_3D_scorername.h5` is located. This can be done using:
```python
-deeplabcut.create_labeled_video_3d(config_path, ['triangulated_file_folder'], start=50, end=250)
+deeplabcut.create_labeled_video_3d(
+ config_path,
+ ["triangulated_file_folder"],
+ start=50,
+ end=250
+)
```
-**TIP:** (see more parameters below) You can set how the axis of the 3D plot on the far right looks by changing the variables ``xlim``, ``ylim``, ``zlim`` and ``view``. Your checkerboard_3d.png image which was created above will show you the axis ranges. Here is an example:
+**TIP:** (see more parameters below) You can set how the axis of the 3D plot on the far right looks by changing the
+variables `xlim`, `ylim`, `zlim` and `view`. Your checkerboard_3d.png image which was created above will show you the
+axis ranges. Here is an example:

 diff --git a/docs/beginner-guides/labeling.md b/docs/beginner-guides/labeling.md
index 7d944137a6..e5c7492722 100644
--- a/docs/beginner-guides/labeling.md
+++ b/docs/beginner-guides/labeling.md
@@ -1,3 +1,4 @@
+(labeling)=
# Labeling GUI
## Selecting Frames to Label
@@ -45,7 +46,9 @@ Alright, you've got your extracted frames ready. Now comes the labeling!
- **Navigate Through Frames:** Use the slider to go from one frame to the next after you're done labeling.
- **Save Progress:** Remember to save your work as you go with **`Command and S`** (or **`Ctrl and S`** on Windows).
-> 💡 **Note:** For a detailed walkthrough on using the Napari labeling GUI, have a look at the [DeepLabCut Napari Guide](https://deeplabcut.github.io/DeepLabCut/docs/napari_GUI.html). Additionally, you can watch our instructional [YouTube video](https://www.youtube.com/watch?v=hsA9IB5r73E) for more insights and tips.
+> 💡 **Note:** For a detailed walkthrough on using the Napari labeling GUI, have a look at the
+[DeepLabCut Napari Guide](napari-gui). Additionally, you can watch our instructional
+[YouTube video](https://www.youtube.com/watch?v=hsA9IB5r73E) for more insights and tips.
### Completing the Set
diff --git a/docs/beginner-guides/manage-project.md b/docs/beginner-guides/manage-project.md
index a53789360f..3f26589ae2 100644
--- a/docs/beginner-guides/manage-project.md
+++ b/docs/beginner-guides/manage-project.md
@@ -41,4 +41,4 @@ A **`Configuration Editor`** window will open, displaying all the configuration
- **Save the Configuration:** Once you're satisfied with the modifications, click **`Save`**. This will store your changes and return you to the main GUI window.
-## Next, head over the beginner guide for [Labeling your data](https://deeplabcut.github.io/DeepLabCut/docs/labelling)
+## Next, head over the beginner guide for [Labeling your data](labeling)
diff --git a/docs/beginner-guides/video-analysis.md b/docs/beginner-guides/video-analysis.md
index 59c976360d..849d8b6638 100644
--- a/docs/beginner-guides/video-analysis.md
+++ b/docs/beginner-guides/video-analysis.md
@@ -16,7 +16,7 @@ After training and evaluating your model, the next step is to apply it to your v
- **Find Results in Your Project Folder:** After analysis, go to your project's video folder.
- **Analysis Files:** Look also for a `.metapickle`, an `.h5`, and possibly a `.csv` file for detailed analysis data.
-- **Review the Plot Poses Subfolder:** This contains visual outputs of the video analysis.
+- **Review the "plot-poses" subfolder:** This contains visual outputs of the video analysis.

diff --git a/docs/convert_maDLC.md b/docs/convert_maDLC.md
index 6671afb1ef..19dd017692 100644
--- a/docs/convert_maDLC.md
+++ b/docs/convert_maDLC.md
@@ -1,10 +1,12 @@
(convert-maDLC)=
-# How to convert a pre-2.2 project for use with DeepLabCut 2.2
+# How to convert a pre-2.2 project for use with DeepLabCut 2.2 or later
diff --git a/docs/beginner-guides/labeling.md b/docs/beginner-guides/labeling.md
index 7d944137a6..e5c7492722 100644
--- a/docs/beginner-guides/labeling.md
+++ b/docs/beginner-guides/labeling.md
@@ -1,3 +1,4 @@
+(labeling)=
# Labeling GUI
## Selecting Frames to Label
@@ -45,7 +46,9 @@ Alright, you've got your extracted frames ready. Now comes the labeling!
- **Navigate Through Frames:** Use the slider to go from one frame to the next after you're done labeling.
- **Save Progress:** Remember to save your work as you go with **`Command and S`** (or **`Ctrl and S`** on Windows).
-> 💡 **Note:** For a detailed walkthrough on using the Napari labeling GUI, have a look at the [DeepLabCut Napari Guide](https://deeplabcut.github.io/DeepLabCut/docs/napari_GUI.html). Additionally, you can watch our instructional [YouTube video](https://www.youtube.com/watch?v=hsA9IB5r73E) for more insights and tips.
+> 💡 **Note:** For a detailed walkthrough on using the Napari labeling GUI, have a look at the
+[DeepLabCut Napari Guide](napari-gui). Additionally, you can watch our instructional
+[YouTube video](https://www.youtube.com/watch?v=hsA9IB5r73E) for more insights and tips.
### Completing the Set
diff --git a/docs/beginner-guides/manage-project.md b/docs/beginner-guides/manage-project.md
index a53789360f..3f26589ae2 100644
--- a/docs/beginner-guides/manage-project.md
+++ b/docs/beginner-guides/manage-project.md
@@ -41,4 +41,4 @@ A **`Configuration Editor`** window will open, displaying all the configuration
- **Save the Configuration:** Once you're satisfied with the modifications, click **`Save`**. This will store your changes and return you to the main GUI window.
-## Next, head over the beginner guide for [Labeling your data](https://deeplabcut.github.io/DeepLabCut/docs/labelling)
+## Next, head over the beginner guide for [Labeling your data](labeling)
diff --git a/docs/beginner-guides/video-analysis.md b/docs/beginner-guides/video-analysis.md
index 59c976360d..849d8b6638 100644
--- a/docs/beginner-guides/video-analysis.md
+++ b/docs/beginner-guides/video-analysis.md
@@ -16,7 +16,7 @@ After training and evaluating your model, the next step is to apply it to your v
- **Find Results in Your Project Folder:** After analysis, go to your project's video folder.
- **Analysis Files:** Look also for a `.metapickle`, an `.h5`, and possibly a `.csv` file for detailed analysis data.
-- **Review the Plot Poses Subfolder:** This contains visual outputs of the video analysis.
+- **Review the "plot-poses" subfolder:** This contains visual outputs of the video analysis.

diff --git a/docs/convert_maDLC.md b/docs/convert_maDLC.md
index 6671afb1ef..19dd017692 100644
--- a/docs/convert_maDLC.md
+++ b/docs/convert_maDLC.md
@@ -1,10 +1,12 @@
(convert-maDLC)=
-# How to convert a pre-2.2 project for use with DeepLabCut 2.2
+# How to convert a pre-2.2 project for use with DeepLabCut 2.2 or later
 -If you have a pre-2.2 project (`labeled-data`) with a **single animal** that you want to use with a multianimal project in DLC 2.2, i.e. use your older data to now train the new multi-task deep neural network, here is what you need to do.
+If you have a pre-2.2 project (`labeled-data`) with a **single animal** that you want to use with a multianimal project
+in DLC 2.2 or later, i.e. use your older data to now train the new multi-task deep neural network, here is what you
+need to do.
(1) We recommend you make a back-up of your project folder.
@@ -14,7 +16,8 @@ If you have a pre-2.2 project (`labeled-data`) with a **single animal** that you
-If you have a pre-2.2 project (`labeled-data`) with a **single animal** that you want to use with a multianimal project in DLC 2.2, i.e. use your older data to now train the new multi-task deep neural network, here is what you need to do.
+If you have a pre-2.2 project (`labeled-data`) with a **single animal** that you want to use with a multianimal project
+in DLC 2.2 or later, i.e. use your older data to now train the new multi-task deep neural network, here is what you
+need to do.
(1) We recommend you make a back-up of your project folder.
@@ -14,7 +16,8 @@ If you have a pre-2.2 project (`labeled-data`) with a **single animal** that you
 -- After `task, scorer, date, project_path` please add the following (i.e. in the image above, you would start adding below line 6) Note, the ordering isn't important but useful to keep consistent with the template:
+- After `task, scorer, date, project_path` please add the following (i.e. in the image above, you would start adding
+below line 6) Note, the ordering isn't important but useful to keep consistent with the template:
```python
multianimalproject: true
@@ -29,9 +32,12 @@ individuals:
- mouse1
```
-- `"uniquebodyparts: []` can stay blank, unless you have other items labeled you want to estimate (consider these as similar to bodyparts in pre-2.2); i.e. corners of a box, etc. All unique bodyparts should not be connected to the multianimal bodyparts in the skeleton you will eventually make. But see "advanced option" below.
+- `"uniquebodyparts: []` can stay blank, unless you have other items labeled you want to estimate (consider these as
+similar to bodyparts in pre-2.2); i.e. corners of a box, etc. All unique bodyparts should not be connected to the
+multianimal bodyparts in the skeleton you will eventually make. See "advanced option" below.
-- Please move your "bodyparts:" to "multianimalbodyparts:" (bodypart names must stay the same!) These are the parts that will always be interconnected fully!
+- Please move your "bodyparts:" to "multianimalbodyparts:" (bodypart names must stay the same!) These are the parts
+that will always be interconnected fully!
```python
multianimalbodyparts:
- snout
@@ -46,20 +52,25 @@ then you can set `bodyparts: MULTI!`
deeplabcut.convert2_maDLC(path_config_file, userfeedback=True)
```
-Now you will see that your data within `labeled-data` are converted to a new format, and the single animal format was saved for you under a new file named `CollectedData_ ...singleanimal.h5` and `.csv` as a back-up!
+Now you will see that your data within `labeled-data` are converted to a new format, and the single animal format was
+saved for you under a new file named `CollectedData_ ...singleanimal.h5` and `.csv` as a back-up!
-(4) We strongly recommend to first run check_labels and verify that the conversion was as expected before creating a multianimal training dataset. For instance, you can load this project `config.yaml` in the Project Manager GUI and check labels then create a multi-animal training set with
+(4) We strongly recommend to first run check_labels and verify that the conversion was as expected before creating a
+multianimal training dataset. For instance, you can load this project `config.yaml` in the Project Manager GUI and
+check labels then create a multi-animal training set with
```python
deeplabcut.create_multianimaltraining_dataset(path_config_file)
```
to begin training.
-**Advanced option:** You can also assign former `bodyparts` to either `uniquebodyparts` or `multianimalbodyparts` (you can even leave some unassigned, which means they will be dropped in the conversion).
+**Advanced option:** You can also assign former `bodyparts` to either `uniquebodyparts` or `multianimalbodyparts`
+(you can even leave some unassigned, which means they will be dropped in the conversion).
Example: Imagine you had a project with the moon and a rocket with two parts labeled:
`bodyparts: [moon, rocket_tip,rocket_bottom]`
-Now you want to use this former project (labeled-data) and work on a new dataset (videos) with one moon but multiple (3) rockets. Then convert it as follows:
+Now you want to use this former project (labeled-data) and work on a new dataset (videos) with one moon but multiple
+(3) rockets. Then convert it as follows:
```
individuals: [rocket1, rocket2, rocket3]
uniquebodyparts: [moon]
diff --git a/docs/course.md b/docs/course.md
index 2ad096995f..b71438cb4f 100644
--- a/docs/course.md
+++ b/docs/course.md
@@ -1,5 +1,10 @@
## DeepLabCut Self-paced Course
+::::{warning}
+This course was designed for DLC 2.
+An updated version for DLC 3 is in the works.
+::::
+
Do you have video of animal behaviors? Step 1: Get Poses ...
-- After `task, scorer, date, project_path` please add the following (i.e. in the image above, you would start adding below line 6) Note, the ordering isn't important but useful to keep consistent with the template:
+- After `task, scorer, date, project_path` please add the following (i.e. in the image above, you would start adding
+below line 6) Note, the ordering isn't important but useful to keep consistent with the template:
```python
multianimalproject: true
@@ -29,9 +32,12 @@ individuals:
- mouse1
```
-- `"uniquebodyparts: []` can stay blank, unless you have other items labeled you want to estimate (consider these as similar to bodyparts in pre-2.2); i.e. corners of a box, etc. All unique bodyparts should not be connected to the multianimal bodyparts in the skeleton you will eventually make. But see "advanced option" below.
+- `"uniquebodyparts: []` can stay blank, unless you have other items labeled you want to estimate (consider these as
+similar to bodyparts in pre-2.2); i.e. corners of a box, etc. All unique bodyparts should not be connected to the
+multianimal bodyparts in the skeleton you will eventually make. See "advanced option" below.
-- Please move your "bodyparts:" to "multianimalbodyparts:" (bodypart names must stay the same!) These are the parts that will always be interconnected fully!
+- Please move your "bodyparts:" to "multianimalbodyparts:" (bodypart names must stay the same!) These are the parts
+that will always be interconnected fully!
```python
multianimalbodyparts:
- snout
@@ -46,20 +52,25 @@ then you can set `bodyparts: MULTI!`
deeplabcut.convert2_maDLC(path_config_file, userfeedback=True)
```
-Now you will see that your data within `labeled-data` are converted to a new format, and the single animal format was saved for you under a new file named `CollectedData_ ...singleanimal.h5` and `.csv` as a back-up!
+Now you will see that your data within `labeled-data` are converted to a new format, and the single animal format was
+saved for you under a new file named `CollectedData_ ...singleanimal.h5` and `.csv` as a back-up!
-(4) We strongly recommend to first run check_labels and verify that the conversion was as expected before creating a multianimal training dataset. For instance, you can load this project `config.yaml` in the Project Manager GUI and check labels then create a multi-animal training set with
+(4) We strongly recommend to first run check_labels and verify that the conversion was as expected before creating a
+multianimal training dataset. For instance, you can load this project `config.yaml` in the Project Manager GUI and
+check labels then create a multi-animal training set with
```python
deeplabcut.create_multianimaltraining_dataset(path_config_file)
```
to begin training.
-**Advanced option:** You can also assign former `bodyparts` to either `uniquebodyparts` or `multianimalbodyparts` (you can even leave some unassigned, which means they will be dropped in the conversion).
+**Advanced option:** You can also assign former `bodyparts` to either `uniquebodyparts` or `multianimalbodyparts`
+(you can even leave some unassigned, which means they will be dropped in the conversion).
Example: Imagine you had a project with the moon and a rocket with two parts labeled:
`bodyparts: [moon, rocket_tip,rocket_bottom]`
-Now you want to use this former project (labeled-data) and work on a new dataset (videos) with one moon but multiple (3) rockets. Then convert it as follows:
+Now you want to use this former project (labeled-data) and work on a new dataset (videos) with one moon but multiple
+(3) rockets. Then convert it as follows:
```
individuals: [rocket1, rocket2, rocket3]
uniquebodyparts: [moon]
diff --git a/docs/course.md b/docs/course.md
index 2ad096995f..b71438cb4f 100644
--- a/docs/course.md
+++ b/docs/course.md
@@ -1,5 +1,10 @@
## DeepLabCut Self-paced Course
+::::{warning}
+This course was designed for DLC 2.
+An updated version for DLC 3 is in the works.
+::::
+
Do you have video of animal behaviors? Step 1: Get Poses ...
 @@ -17,7 +22,7 @@ We expect it to take *roughly* 1-2 weeks to get through if you do it rigorously.
## Installation:
You need Python and DeepLabCut installed!
-- [See these "beginner docs" for help!](https://deeplabcut.github.io/DeepLabCut/docs/beginners-guide.html)
+- [See these "beginner docs" for help!](beginners-guide)
- **WATCH:** overview of conda: [Python Tutorial: Anaconda - Installation and Using Conda](https://www.youtube.com/watch?v=YJC6ldI3hWk)
@@ -32,10 +37,8 @@ You need Python and DeepLabCut installed!
- **Learning:** learning and teaching signal processing, and overview from Prof. Demba Ba [talk at JupyterCon](https://www.youtube.com/watch?v=ywz-LLYwkQQ)
-- **Learning:** Watch a talk from Alexander Mathis (a lead DeepLabCut developer) [talk about DeepLabCut!](https://www.youtube.com/watch?v=ZjWPHM0sL4E)
-
- **DEMO:** Can I DEMO DEEPLABCUT (DLC) quickly?
- - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
+ - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
- AND follow along with me: [Video Tutorial!](https://www.youtube.com/watch?v=DRT-Cq2vdWs)
@@ -55,9 +58,9 @@ You need Python and DeepLabCut installed!
**What you need:** any videos where you can see the animals/objects, etc.
You can use our demo videos, grab some from the internet, or use whatever older data you have. Any camera, color/monochrome, etc will work. Find diverse videos, and label what you want to track well :)
-- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo :smile:
+- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo 😊
- - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/master/part1-labeling.pdf)
+ - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part1-labeling.pdf)
- **READ ME PLEASE:** [DeepLabCut, the science](https://rdcu.be/4Rep)
- **READ ME PLEASE:** [DeepLabCut, the user guide](https://rdcu.be/bHpHN)
- **WATCH:** Video tutorial 1: [using the Project Manager GUI](https://www.youtube.com/watch?v=KcXogR-p5Ak)
@@ -71,12 +74,12 @@ You can use our demo videos, grab some from the internet, or use whatever older
### **Module 2: Neural Networks**
- - **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/master/part2-network.pdf)
+ - **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part2-network.pdf)
- **READ ME PLEASE:** [What are convolutional neural networks?](https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
- **READ ME PLEASE:** Here is a new paper from us describing challenges in robust pose estimation, why PRE-TRAINING really matters - which was our major scientific contribution to low-data input pose-estimation - and it describes new networks that are available to you. [Pretraining boosts out-of-domain robustness for pose estimation](https://paperswithcode.com/paper/pretraining-boosts-out-of-domain-robustness)
- - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/ (link to [ppt from Dr. Fei-Fei Li](http://www.image-net.org/papers/ImageNet_2010.ppt))
+ - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/
- **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
@@ -84,18 +87,18 @@ You can use our demo videos, grab some from the internet, or use whatever older
@@ -17,7 +22,7 @@ We expect it to take *roughly* 1-2 weeks to get through if you do it rigorously.
## Installation:
You need Python and DeepLabCut installed!
-- [See these "beginner docs" for help!](https://deeplabcut.github.io/DeepLabCut/docs/beginners-guide.html)
+- [See these "beginner docs" for help!](beginners-guide)
- **WATCH:** overview of conda: [Python Tutorial: Anaconda - Installation and Using Conda](https://www.youtube.com/watch?v=YJC6ldI3hWk)
@@ -32,10 +37,8 @@ You need Python and DeepLabCut installed!
- **Learning:** learning and teaching signal processing, and overview from Prof. Demba Ba [talk at JupyterCon](https://www.youtube.com/watch?v=ywz-LLYwkQQ)
-- **Learning:** Watch a talk from Alexander Mathis (a lead DeepLabCut developer) [talk about DeepLabCut!](https://www.youtube.com/watch?v=ZjWPHM0sL4E)
-
- **DEMO:** Can I DEMO DEEPLABCUT (DLC) quickly?
- - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
+ - Yes: [you can click through this DEMO notebook](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DEMO_mouse_openfield.ipynb)
- AND follow along with me: [Video Tutorial!](https://www.youtube.com/watch?v=DRT-Cq2vdWs)
@@ -55,9 +58,9 @@ You need Python and DeepLabCut installed!
**What you need:** any videos where you can see the animals/objects, etc.
You can use our demo videos, grab some from the internet, or use whatever older data you have. Any camera, color/monochrome, etc will work. Find diverse videos, and label what you want to track well :)
-- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo :smile:
+- IF YOU ARE PART OF THE COURSE: you will be contributing to the DLC Model Zoo 😊
- - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/master/part1-labeling.pdf)
+ - **Slides:** [Overview of starting new projects](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part1-labeling.pdf)
- **READ ME PLEASE:** [DeepLabCut, the science](https://rdcu.be/4Rep)
- **READ ME PLEASE:** [DeepLabCut, the user guide](https://rdcu.be/bHpHN)
- **WATCH:** Video tutorial 1: [using the Project Manager GUI](https://www.youtube.com/watch?v=KcXogR-p5Ak)
@@ -71,12 +74,12 @@ You can use our demo videos, grab some from the internet, or use whatever older
### **Module 2: Neural Networks**
- - **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/master/part2-network.pdf)
+ - **Slides:** [Overview of creating training and test data, and training networks](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part2-network.pdf)
- **READ ME PLEASE:** [What are convolutional neural networks?](https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
- **READ ME PLEASE:** Here is a new paper from us describing challenges in robust pose estimation, why PRE-TRAINING really matters - which was our major scientific contribution to low-data input pose-estimation - and it describes new networks that are available to you. [Pretraining boosts out-of-domain robustness for pose estimation](https://paperswithcode.com/paper/pretraining-boosts-out-of-domain-robustness)
- - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/ (link to [ppt from Dr. Fei-Fei Li](http://www.image-net.org/papers/ImageNet_2010.ppt))
+ - **MORE DETAILS:** ImageNet: check out the original paper and dataset: http://www.image-net.org/
- **REVIEW PAPER:** [A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives](https://www.sciencedirect.com/science/article/pii/S0896627320307170)
@@ -84,18 +87,18 @@ You can use our demo videos, grab some from the internet, or use whatever older
 Before you create a training/test set, please read/watch:
- - **More information:** [Which types neural networks are available, and what should I use?](https://github.com/AlexEMG/DeepLabCut/wiki/What-neural-network-should-I-use%3F)
+ - **More information:** [Which types neural networks are available, and what should I use?](https://github.com/DeepLabCut/DeepLabCut/wiki/What-neural-network-should-I-use%3F-(Trade-offs,-speed-performance,-and-considerations))
- **WATCH:** Video tutorial 1: [How to test different networks in a controlled way](https://www.youtube.com/watch?v=WXCVr6xAcCA)
- Now, decide what model(s) you want to test.
- IF you want to train on your CPU, then run the step `create_training_dataset`, in the GUI etc. on your own computer.
- - IF you want to use GPUs on google colab, [**(1)** watch this FIRST/follow along here!](https://www.youtube.com/watch?v=qJGs8nxx80A) **(2)** move your whole project folder to Google Drive, and then [**use this notebook**](https://github.com/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb)
+ - IF you want to use GPUs on google colab, [**(1)** watch this FIRST/follow along here!](https://www.youtube.com/watch?v=qJGs8nxx80A) **(2)** move your whole project folder to Google Drive, and then [**use this notebook**](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb)
**MODULE 2 webinar**: https://youtu.be/ILsuC4icBU0
### **Module 3: Evalution of network performance**
- - **Slides** [Evalute your network](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/master/part3-analysis.pdf)
+ - **Slides** [Evalute your network](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part3-analysis.pdf)
- **WATCH:** [Evaluate the network in ipython](https://www.youtube.com/watch?v=bgfnz1wtlpo)
- why evaluation matters; how to benchmark; analyzing a video and using scoremaps, conf. readouts, etc.
diff --git a/docs/deeplabcutlive.md b/docs/deeplabcutlive.md
index f8784b651d..f1edbff3cc 100644
--- a/docs/deeplabcutlive.md
+++ b/docs/deeplabcutlive.md
@@ -1,3 +1,4 @@
+(deeplabcut-live)=
# DeepLabCut-Live!
We provide two additional pip packages that allow you to record and stream camera data and run DeeplabCut models in real-time.
diff --git a/docs/gui/PROJECT_GUI.md b/docs/gui/PROJECT_GUI.md
index 106e7eca5c..e0883c324e 100644
--- a/docs/gui/PROJECT_GUI.md
+++ b/docs/gui/PROJECT_GUI.md
@@ -10,7 +10,7 @@ As some users may be more comfortable working with an interactive interface, we
(1) Install DeepLabCut using the simple-install with Anaconda found [here!](how-to-install)*.
Now you have DeepLabCut installed, but if you want to update it, either follow the prompt in the GUI which will ask you to upgrade when a new version is available, or just go into your env (activate DEEPLABCUT) then run:
-` pip install 'deeplabcut[gui,modelzoo]'` *but please see [full install guide](https://deeplabcut.github.io/DeepLabCut/docs/installation.html)!
+` pip install 'deeplabcut[gui,modelzoo]'` *but please see [full install guide](how-to-install)!
(2) Open the terminal and run: `python -m deeplabcut`
@@ -23,15 +23,14 @@ Now you have DeepLabCut installed, but if you want to update it, either follow t
Start at the Project Management Tab and work your way through the tabs to built your customized model and deploy it on new data.
We recommend to keep the terminal visible (as well as the GUI) so you can see the ongoing processes as you step through your project, or any errors that might arise.
-- For specific napari-based labeling features, see the ["napari gui" docs](https://deeplabcut.github.io/DeepLabCut/docs/napari_GUI.html#usage).
+- For specific napari-based labeling features, see the ["napari gui" docs](napari-gui-usage).
- To change from dark to light mode, set appearance at the top:
Before you create a training/test set, please read/watch:
- - **More information:** [Which types neural networks are available, and what should I use?](https://github.com/AlexEMG/DeepLabCut/wiki/What-neural-network-should-I-use%3F)
+ - **More information:** [Which types neural networks are available, and what should I use?](https://github.com/DeepLabCut/DeepLabCut/wiki/What-neural-network-should-I-use%3F-(Trade-offs,-speed-performance,-and-considerations))
- **WATCH:** Video tutorial 1: [How to test different networks in a controlled way](https://www.youtube.com/watch?v=WXCVr6xAcCA)
- Now, decide what model(s) you want to test.
- IF you want to train on your CPU, then run the step `create_training_dataset`, in the GUI etc. on your own computer.
- - IF you want to use GPUs on google colab, [**(1)** watch this FIRST/follow along here!](https://www.youtube.com/watch?v=qJGs8nxx80A) **(2)** move your whole project folder to Google Drive, and then [**use this notebook**](https://github.com/DeepLabCut/DeepLabCut/blob/master/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb)
+ - IF you want to use GPUs on google colab, [**(1)** watch this FIRST/follow along here!](https://www.youtube.com/watch?v=qJGs8nxx80A) **(2)** move your whole project folder to Google Drive, and then [**use this notebook**](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_YOURDATA_TrainNetwork_VideoAnalysis.ipynb)
**MODULE 2 webinar**: https://youtu.be/ILsuC4icBU0
### **Module 3: Evalution of network performance**
- - **Slides** [Evalute your network](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/master/part3-analysis.pdf)
+ - **Slides** [Evalute your network](https://github.com/DeepLabCut/DeepLabCut-Workshop-Materials/blob/main/part3-analysis.pdf)
- **WATCH:** [Evaluate the network in ipython](https://www.youtube.com/watch?v=bgfnz1wtlpo)
- why evaluation matters; how to benchmark; analyzing a video and using scoremaps, conf. readouts, etc.
diff --git a/docs/deeplabcutlive.md b/docs/deeplabcutlive.md
index f8784b651d..f1edbff3cc 100644
--- a/docs/deeplabcutlive.md
+++ b/docs/deeplabcutlive.md
@@ -1,3 +1,4 @@
+(deeplabcut-live)=
# DeepLabCut-Live!
We provide two additional pip packages that allow you to record and stream camera data and run DeeplabCut models in real-time.
diff --git a/docs/gui/PROJECT_GUI.md b/docs/gui/PROJECT_GUI.md
index 106e7eca5c..e0883c324e 100644
--- a/docs/gui/PROJECT_GUI.md
+++ b/docs/gui/PROJECT_GUI.md
@@ -10,7 +10,7 @@ As some users may be more comfortable working with an interactive interface, we
(1) Install DeepLabCut using the simple-install with Anaconda found [here!](how-to-install)*.
Now you have DeepLabCut installed, but if you want to update it, either follow the prompt in the GUI which will ask you to upgrade when a new version is available, or just go into your env (activate DEEPLABCUT) then run:
-` pip install 'deeplabcut[gui,modelzoo]'` *but please see [full install guide](https://deeplabcut.github.io/DeepLabCut/docs/installation.html)!
+` pip install 'deeplabcut[gui,modelzoo]'` *but please see [full install guide](how-to-install)!
(2) Open the terminal and run: `python -m deeplabcut`
@@ -23,15 +23,14 @@ Now you have DeepLabCut installed, but if you want to update it, either follow t
Start at the Project Management Tab and work your way through the tabs to built your customized model and deploy it on new data.
We recommend to keep the terminal visible (as well as the GUI) so you can see the ongoing processes as you step through your project, or any errors that might arise.
-- For specific napari-based labeling features, see the ["napari gui" docs](https://deeplabcut.github.io/DeepLabCut/docs/napari_GUI.html#usage).
+- For specific napari-based labeling features, see the ["napari gui" docs](napari-gui-usage).
- To change from dark to light mode, set appearance at the top:

 +
+ ```{Hint}
**Labeling Pitfalls: How Corruptions Affect Performance**
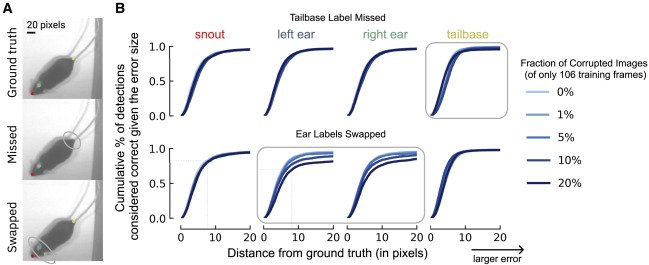
-(A) Illustration of two types of labeling errors. Top is ground truth, middle is missing a label at the tailbase, and bottom is if the labeler swapped the ear identity (left to right, etc.). (B) Using a small training dataset of 106 frames, how do the corruptions in (A) affect the percent of correct keypoints (PCK) on the test set as the distance to ground truth increases from 0 pixels (perfect prediction) to 20 pixels (larger error)? The x axis denotes the difference in the ground truth to the predicted location (RMSE in pixels), whereas the y axis is the fraction of frames considered accurate (e.g., z80% of frames fall within 9 pixels, even on this small training dataset, for points that are not corrupted, whereas for swapped points this falls to z65%). The fraction of the dataset that is corrupted affects this value. Shown is when missing the tailbase label (top) or swapping the ears in 1%, 5%, 10%, and 20% of frames (of 106 labeled training images). Swapping versus missing labels has a more notable adverse effect on network performance.
+(A) Illustration of two types of labeling errors. Top is ground truth, middle is missing a label at the tailbase, and
+bottom is if the labeler swapped the ear identity (left to right, etc.). (B) Using a small training dataset of 106
+frames, how do the corruptions in (A) affect the percent of correct keypoints (PCK) on the test set as the distance
+to ground truth increases from 0 pixels (perfect prediction) to 20 pixels (larger error)? The x axis denotes the
+difference in the ground truth to the predicted location (RMSE in pixels), whereas the y axis is the fraction of
+frames considered accurate (e.g., z80% of frames fall within 9 pixels, even on this small training dataset, for
+points that are not corrupted, whereas for swapped points this falls to z65%). The fraction of the dataset that is
+corrupted affects this value. Shown is when missing the tailbase label (top) or swapping the ears in 1%, 5%, 10%,
+and 20% of frames (of 106 labeled training images). Swapping versus missing labels has a more notable adverse effect
+on network performance.
```
-The DeepLabCut toolbox supports **active learning** by extracting outlier frames be several methods and allowing the user to correct the frames, then retrain the model. See the [Nature Protocols paper](https://www.nature.com/articles/s41596-019-0176-0) for the detailed steps, or in the docs, [here](https://deeplabcut.github.io/DeepLabCut/docs/standardDeepLabCut_UserGuide.html#m-optional-active-learning-network-refinement-extract-outlier-frames).
+The DeepLabCut toolbox supports **active learning** by extracting outlier frames be several methods and allowing the
+user to correct the frames, then retrain the model. See the
+[Nature Protocols paper](https://www.nature.com/articles/s41596-019-0176-0) for the detailed steps, or in the docs,
+[here](active-learning).
-To facilitate this process, here we propose a new way to detect 'outlier frames', which is planned to be released in ~Sept 2022. Your contributions and suggestions are welcomed, so test the [PR](https://github.com/DeepLabCut/napari-deeplabcut/pull/38) and give us feedback!
+To facilitate this process, here we propose a new way to detect 'outlier frames'.
+Your contributions and suggestions are welcomed, so test the
+[PR](https://github.com/DeepLabCut/napari-deeplabcut/pull/38) and give us feedback!
-This #cookbook recipe aims to show a usecase of **clustering in napari** and is contributed by 2022 DLC AI Resident [Sabrina Benas](https://twitter.com/Sabrineiitor) 💜.
+This #cookbook recipe aims to show a usecase of **clustering in napari** and is contributed by 2022 DLC AI Resident
+[Sabrina Benas](https://twitter.com/Sabrineiitor) 💜.
## Detect Outliers to Refine Labels
### Open `napari` and the `DeepLabCut plugin`
- - Then open your `CollectedData_
```{Hint}
**Labeling Pitfalls: How Corruptions Affect Performance**
-(A) Illustration of two types of labeling errors. Top is ground truth, middle is missing a label at the tailbase, and bottom is if the labeler swapped the ear identity (left to right, etc.). (B) Using a small training dataset of 106 frames, how do the corruptions in (A) affect the percent of correct keypoints (PCK) on the test set as the distance to ground truth increases from 0 pixels (perfect prediction) to 20 pixels (larger error)? The x axis denotes the difference in the ground truth to the predicted location (RMSE in pixels), whereas the y axis is the fraction of frames considered accurate (e.g., z80% of frames fall within 9 pixels, even on this small training dataset, for points that are not corrupted, whereas for swapped points this falls to z65%). The fraction of the dataset that is corrupted affects this value. Shown is when missing the tailbase label (top) or swapping the ears in 1%, 5%, 10%, and 20% of frames (of 106 labeled training images). Swapping versus missing labels has a more notable adverse effect on network performance.
+(A) Illustration of two types of labeling errors. Top is ground truth, middle is missing a label at the tailbase, and
+bottom is if the labeler swapped the ear identity (left to right, etc.). (B) Using a small training dataset of 106
+frames, how do the corruptions in (A) affect the percent of correct keypoints (PCK) on the test set as the distance
+to ground truth increases from 0 pixels (perfect prediction) to 20 pixels (larger error)? The x axis denotes the
+difference in the ground truth to the predicted location (RMSE in pixels), whereas the y axis is the fraction of
+frames considered accurate (e.g., z80% of frames fall within 9 pixels, even on this small training dataset, for
+points that are not corrupted, whereas for swapped points this falls to z65%). The fraction of the dataset that is
+corrupted affects this value. Shown is when missing the tailbase label (top) or swapping the ears in 1%, 5%, 10%,
+and 20% of frames (of 106 labeled training images). Swapping versus missing labels has a more notable adverse effect
+on network performance.
```
-The DeepLabCut toolbox supports **active learning** by extracting outlier frames be several methods and allowing the user to correct the frames, then retrain the model. See the [Nature Protocols paper](https://www.nature.com/articles/s41596-019-0176-0) for the detailed steps, or in the docs, [here](https://deeplabcut.github.io/DeepLabCut/docs/standardDeepLabCut_UserGuide.html#m-optional-active-learning-network-refinement-extract-outlier-frames).
+The DeepLabCut toolbox supports **active learning** by extracting outlier frames be several methods and allowing the
+user to correct the frames, then retrain the model. See the
+[Nature Protocols paper](https://www.nature.com/articles/s41596-019-0176-0) for the detailed steps, or in the docs,
+[here](active-learning).
-To facilitate this process, here we propose a new way to detect 'outlier frames', which is planned to be released in ~Sept 2022. Your contributions and suggestions are welcomed, so test the [PR](https://github.com/DeepLabCut/napari-deeplabcut/pull/38) and give us feedback!
+To facilitate this process, here we propose a new way to detect 'outlier frames'.
+Your contributions and suggestions are welcomed, so test the
+[PR](https://github.com/DeepLabCut/napari-deeplabcut/pull/38) and give us feedback!
-This #cookbook recipe aims to show a usecase of **clustering in napari** and is contributed by 2022 DLC AI Resident [Sabrina Benas](https://twitter.com/Sabrineiitor) 💜.
+This #cookbook recipe aims to show a usecase of **clustering in napari** and is contributed by 2022 DLC AI Resident
+[Sabrina Benas](https://twitter.com/Sabrineiitor) 💜.
## Detect Outliers to Refine Labels
### Open `napari` and the `DeepLabCut plugin`
- - Then open your `CollectedData_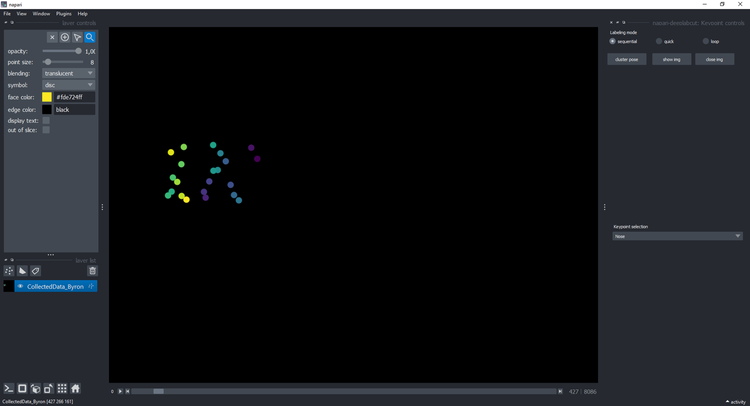 +
+ ### Clustering
-- Click on the button `cluster` and wait a few seconds until it displays a new layer with the cluster:
+Click on the button `cluster` and wait a few seconds until it displays a new layer with the cluster:
-
### Clustering
-- Click on the button `cluster` and wait a few seconds until it displays a new layer with the cluster:
+Click on the button `cluster` and wait a few seconds until it displays a new layer with the cluster:
- 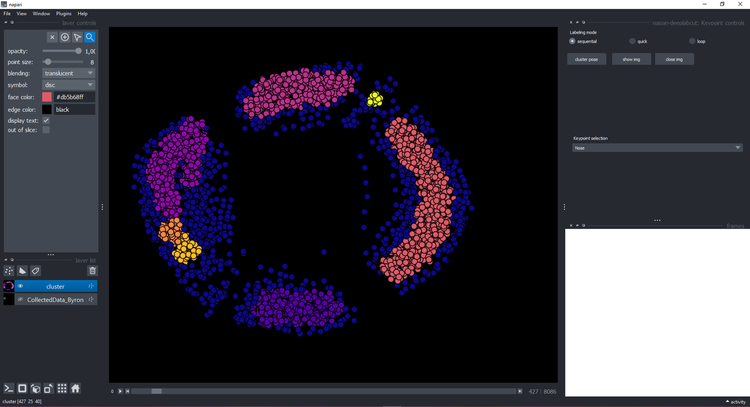 +
+ You can click on a point and see the image on the right with the keypoints:
-
You can click on a point and see the image on the right with the keypoints:
- 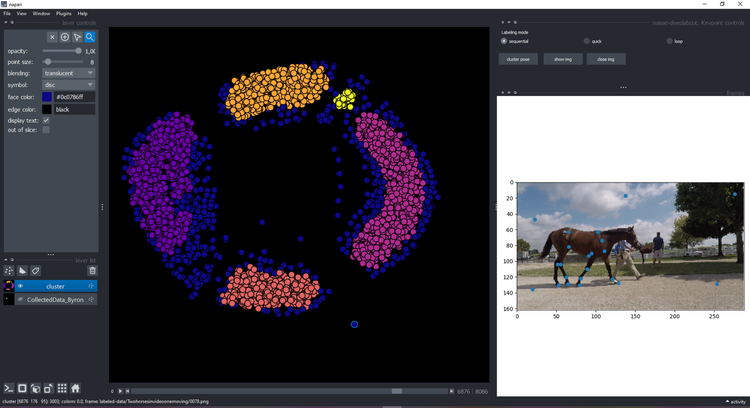 +
+ ### Visualize & refine
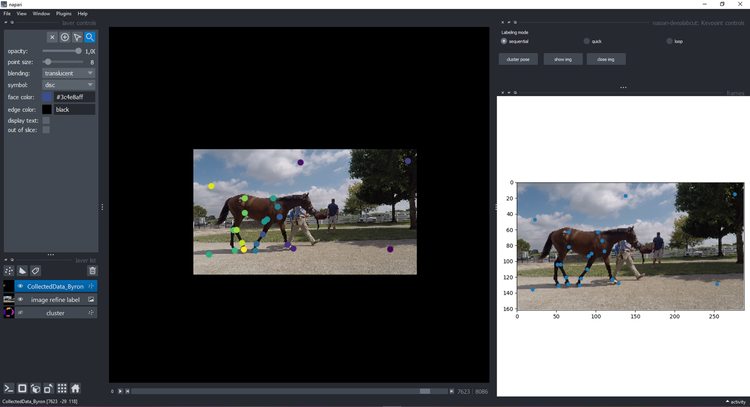
-If you decided to refine that frame (we moved the points to make outliers obvious), click `show img` and refine them using the plugin features and instructions:
+If you decided to refine that frame (we moved the points to make outliers obvious), click `show img` and refine them
+using the plugin features and instructions:
-
### Visualize & refine
-If you decided to refine that frame (we moved the points to make outliers obvious), click `show img` and refine them using the plugin features and instructions:
+If you decided to refine that frame (we moved the points to make outliers obvious), click `show img` and refine them
+using the plugin features and instructions:
-  +
+ - ```{Attention}
- When you're done, you need to click `ctl-s` to save it.
+```{Attention}
+When you're done, you need to click `ctl-s` to save it.
```
-- You can go back to the cluster layer by clicking on `close img` and refine another image. Reminder, when you're done editing you need to click `ctl-s` to save your work. And now you can take the updated `CollectedData` file, create and **new training shuffle**, and train the network! Read more about how to [create a training dataset](https://deeplabcut.github.io/DeepLabCut/docs/standardDeepLabCut_UserGuide.html#f-create-training-dataset-s).
+You can go back to the cluster layer by clicking on `close img` and refine another image. Reminder, when you're done
+editing you need to click `ctl-s` to save your work. And now you can take the updated `CollectedData` file, create
+and **new training shuffle**, and train the network! Read more about how to
+[create a training dataset](create-training-dataset).
```{hint}
-If you want to change the clustering method, you can modify the file [kmeans.py](https://github.com/DeepLabCutAIResidency/napari-deeplabcut/blob/cluster1/src/napari_deeplabcut/kmeans.py)
+If you want to change the clustering method, you can modify the file
+[kmeans.py](https://github.com/DeepLabCutAIResidency/napari-deeplabcut/blob/cluster1/src/napari_deeplabcut/kmeans.py)
+```
::::{important}
-You have to keep the way the file is opened (pandas dataframe) and the output has to be the cluster points, the points colors in the cluster colors and the frame names (in this order).
+You have to keep the way the file is opened (pandas dataframe) and the output has to be the cluster points, the points
+colors in the cluster colors and the frame names (in this order).
::::
```
diff --git a/docs/recipes/DLCMethods.md b/docs/recipes/DLCMethods.md
index b2f7a5a7af..74d5ec4c65 100644
--- a/docs/recipes/DLCMethods.md
+++ b/docs/recipes/DLCMethods.md
@@ -2,7 +2,13 @@
**Pose estimation using DeepLabCut**
-For body part tracking we used DeepLabCut (version 2.X.X) [Mathis et al, 2018, Nath et al, 2019]. Specifically, we labeled X number of frames taken from X videos/animals (then X% was used for training (default is 95%). We used a X-based neural network (i.e., X = ResNet-50, ResNet-101, MobileNetV2-0.35, MobileNetV2-0.5, MobileNetV2-0.75, MobileNetV2-1, EfficientNet ..X, dlcrnet_ms5, etc.)*** with default parameters* for X number of training iterations. We validated with X number of shuffles, and found the test error was: X pixels, train: X pixels (image size was X by X). We then used a p-cutoff of X (i.e. 0.9) to condition the X,Y coordinates for future analysis. This network was then used to analyze videos from similar experimental settings.
+For body part tracking we used DeepLabCut (version 3.X.X) [Mathis et al, 2018, Nath et al, 2019]. Specifically, we
+labeled X number of frames taken from X videos/animals (then X% was used for training (default is 95%). We used a
+X-based neural network (i.e., X = ResNet-50, ResNet-101, MobileNetV2-0.35, MobileNetV2-0.5, MobileNetV2-0.75,
+MobileNetV2-1, EfficientNet ..X, dlcrnet_ms5, cspnext_s, dekr_w32, rtmpose_s, etc.)*** with default parameters* for X
+number of training iterations. We validated with X number of shuffles, and found the test error was: X pixels, train:
+X pixels (image size was X by X). We then used a p-cutoff of X (i.e. 0.9) to condition the X,Y coordinates for future
+analysis. This network was then used to analyze videos from similar experimental settings.
*If any defaults were changed in *`pose_config.yaml`*, mention them.
@@ -43,4 +49,5 @@ If you use ResNets, consider citing Insafutdinov et al 2016 & He et al 2016. If
> 770–778 (2016). URL https://arxiv.org/abs/
> 1512.03385.
-We also have the network graphic freely available on SciDraw.io if you'd like to use it! https://scidraw.io/drawing/290. If you use our DLC logo, please include the TM symbol, thank you!
+We also have the network graphic freely available on SciDraw.io if you'd like to use it! https://scidraw.io/drawing/290.
+If you use our DLC logo, please include the TM symbol, thank you!
diff --git a/docs/recipes/MegaDetectorDLCLive.md b/docs/recipes/MegaDetectorDLCLive.md
index 20e35a38fc..8af6f59d1b 100644
--- a/docs/recipes/MegaDetectorDLCLive.md
+++ b/docs/recipes/MegaDetectorDLCLive.md
@@ -14,7 +14,9 @@ MegaDetector detects an animal and generates a bounding box around the animal. T
## DeepLabCut-Live
-DeepLabCut-Live! is a real-time package for running DeepLabCut. However, you can also use it as a lighter-weight package for running DeeplabCut even if you don't need real-time. It's very useful to use in HPC or servers, or in Apps, as we do here. To read more, check out the [docs](https://deeplabcut.github.io/DeepLabCut/docs/deeplabcutlive.html).
+DeepLabCut-Live! is a real-time package for running DeepLabCut. However, you can also use it as a lighter-weight
+package for running DeeplabCut even if you don't need real-time. It's very useful to use in HPC or servers, or in Apps,
+as we do here. To read more, check out the [docs](deeplabcut-live).
### MegaDetector meets DeepLabCut
diff --git a/docs/recipes/OpenVINO.md b/docs/recipes/OpenVINO.md
index 2033045ce8..78ea18e82f 100644
--- a/docs/recipes/OpenVINO.md
+++ b/docs/recipes/OpenVINO.md
@@ -1,10 +1,15 @@
# Intel OpenVINO backend
+::::{warning}
+This feature is currently implemented for TensorFlow-based models only.
+::::
+
DeepLabCut provides an option to run deep learning model with [OpenVINO](https://github.com/openvinotoolkit/openvino) backend.
-To enable OpenVINO in your pipeline, use `use_openvino` flag of `analyze_videos` method with one of string values indicating device:
-* "CPU" - Use CPU. This is a default value.
-* "GPU" - Use iGPU (requires OpenCL to be installed). First launch might take some time for kernels initialization.
-* "MULTI:CPU,GPU" - Use CPU and GPU simultaneously. In most cases this option provides the best efficiency.
+To enable OpenVINO in your pipeline, use `use_openvino` flag of `analyze_videos` method with one of string values
+indicating device:
+* ```"CPU"``` - Use CPU. This is a default value.
+* ```"GPU"``` - Use GPU (requires OpenCL to be installed). First launch might take some time for kernels initialization.
+* ```"MULTI:CPU,GPU"``` - Use CPU and GPU simultaneously. In most cases this option provides the best efficiency.
```python
def analyze_videos(
diff --git a/docs/recipes/UsingModelZooPupil.md b/docs/recipes/UsingModelZooPupil.md
index 2aac4a4d00..7ea4107d75 100644
--- a/docs/recipes/UsingModelZooPupil.md
+++ b/docs/recipes/UsingModelZooPupil.md
@@ -1,17 +1,25 @@
# Using ModelZoo models on your own datasets
-
- ```{Attention}
- When you're done, you need to click `ctl-s` to save it.
+```{Attention}
+When you're done, you need to click `ctl-s` to save it.
```
-- You can go back to the cluster layer by clicking on `close img` and refine another image. Reminder, when you're done editing you need to click `ctl-s` to save your work. And now you can take the updated `CollectedData` file, create and **new training shuffle**, and train the network! Read more about how to [create a training dataset](https://deeplabcut.github.io/DeepLabCut/docs/standardDeepLabCut_UserGuide.html#f-create-training-dataset-s).
+You can go back to the cluster layer by clicking on `close img` and refine another image. Reminder, when you're done
+editing you need to click `ctl-s` to save your work. And now you can take the updated `CollectedData` file, create
+and **new training shuffle**, and train the network! Read more about how to
+[create a training dataset](create-training-dataset).
```{hint}
-If you want to change the clustering method, you can modify the file [kmeans.py](https://github.com/DeepLabCutAIResidency/napari-deeplabcut/blob/cluster1/src/napari_deeplabcut/kmeans.py)
+If you want to change the clustering method, you can modify the file
+[kmeans.py](https://github.com/DeepLabCutAIResidency/napari-deeplabcut/blob/cluster1/src/napari_deeplabcut/kmeans.py)
+```
::::{important}
-You have to keep the way the file is opened (pandas dataframe) and the output has to be the cluster points, the points colors in the cluster colors and the frame names (in this order).
+You have to keep the way the file is opened (pandas dataframe) and the output has to be the cluster points, the points
+colors in the cluster colors and the frame names (in this order).
::::
```
diff --git a/docs/recipes/DLCMethods.md b/docs/recipes/DLCMethods.md
index b2f7a5a7af..74d5ec4c65 100644
--- a/docs/recipes/DLCMethods.md
+++ b/docs/recipes/DLCMethods.md
@@ -2,7 +2,13 @@
**Pose estimation using DeepLabCut**
-For body part tracking we used DeepLabCut (version 2.X.X) [Mathis et al, 2018, Nath et al, 2019]. Specifically, we labeled X number of frames taken from X videos/animals (then X% was used for training (default is 95%). We used a X-based neural network (i.e., X = ResNet-50, ResNet-101, MobileNetV2-0.35, MobileNetV2-0.5, MobileNetV2-0.75, MobileNetV2-1, EfficientNet ..X, dlcrnet_ms5, etc.)*** with default parameters* for X number of training iterations. We validated with X number of shuffles, and found the test error was: X pixels, train: X pixels (image size was X by X). We then used a p-cutoff of X (i.e. 0.9) to condition the X,Y coordinates for future analysis. This network was then used to analyze videos from similar experimental settings.
+For body part tracking we used DeepLabCut (version 3.X.X) [Mathis et al, 2018, Nath et al, 2019]. Specifically, we
+labeled X number of frames taken from X videos/animals (then X% was used for training (default is 95%). We used a
+X-based neural network (i.e., X = ResNet-50, ResNet-101, MobileNetV2-0.35, MobileNetV2-0.5, MobileNetV2-0.75,
+MobileNetV2-1, EfficientNet ..X, dlcrnet_ms5, cspnext_s, dekr_w32, rtmpose_s, etc.)*** with default parameters* for X
+number of training iterations. We validated with X number of shuffles, and found the test error was: X pixels, train:
+X pixels (image size was X by X). We then used a p-cutoff of X (i.e. 0.9) to condition the X,Y coordinates for future
+analysis. This network was then used to analyze videos from similar experimental settings.
*If any defaults were changed in *`pose_config.yaml`*, mention them.
@@ -43,4 +49,5 @@ If you use ResNets, consider citing Insafutdinov et al 2016 & He et al 2016. If
> 770–778 (2016). URL https://arxiv.org/abs/
> 1512.03385.
-We also have the network graphic freely available on SciDraw.io if you'd like to use it! https://scidraw.io/drawing/290. If you use our DLC logo, please include the TM symbol, thank you!
+We also have the network graphic freely available on SciDraw.io if you'd like to use it! https://scidraw.io/drawing/290.
+If you use our DLC logo, please include the TM symbol, thank you!
diff --git a/docs/recipes/MegaDetectorDLCLive.md b/docs/recipes/MegaDetectorDLCLive.md
index 20e35a38fc..8af6f59d1b 100644
--- a/docs/recipes/MegaDetectorDLCLive.md
+++ b/docs/recipes/MegaDetectorDLCLive.md
@@ -14,7 +14,9 @@ MegaDetector detects an animal and generates a bounding box around the animal. T
## DeepLabCut-Live
-DeepLabCut-Live! is a real-time package for running DeepLabCut. However, you can also use it as a lighter-weight package for running DeeplabCut even if you don't need real-time. It's very useful to use in HPC or servers, or in Apps, as we do here. To read more, check out the [docs](https://deeplabcut.github.io/DeepLabCut/docs/deeplabcutlive.html).
+DeepLabCut-Live! is a real-time package for running DeepLabCut. However, you can also use it as a lighter-weight
+package for running DeeplabCut even if you don't need real-time. It's very useful to use in HPC or servers, or in Apps,
+as we do here. To read more, check out the [docs](deeplabcut-live).
### MegaDetector meets DeepLabCut
diff --git a/docs/recipes/OpenVINO.md b/docs/recipes/OpenVINO.md
index 2033045ce8..78ea18e82f 100644
--- a/docs/recipes/OpenVINO.md
+++ b/docs/recipes/OpenVINO.md
@@ -1,10 +1,15 @@
# Intel OpenVINO backend
+::::{warning}
+This feature is currently implemented for TensorFlow-based models only.
+::::
+
DeepLabCut provides an option to run deep learning model with [OpenVINO](https://github.com/openvinotoolkit/openvino) backend.
-To enable OpenVINO in your pipeline, use `use_openvino` flag of `analyze_videos` method with one of string values indicating device:
-* "CPU" - Use CPU. This is a default value.
-* "GPU" - Use iGPU (requires OpenCL to be installed). First launch might take some time for kernels initialization.
-* "MULTI:CPU,GPU" - Use CPU and GPU simultaneously. In most cases this option provides the best efficiency.
+To enable OpenVINO in your pipeline, use `use_openvino` flag of `analyze_videos` method with one of string values
+indicating device:
+* ```"CPU"``` - Use CPU. This is a default value.
+* ```"GPU"``` - Use GPU (requires OpenCL to be installed). First launch might take some time for kernels initialization.
+* ```"MULTI:CPU,GPU"``` - Use CPU and GPU simultaneously. In most cases this option provides the best efficiency.
```python
def analyze_videos(
diff --git a/docs/recipes/UsingModelZooPupil.md b/docs/recipes/UsingModelZooPupil.md
index 2aac4a4d00..7ea4107d75 100644
--- a/docs/recipes/UsingModelZooPupil.md
+++ b/docs/recipes/UsingModelZooPupil.md
@@ -1,17 +1,25 @@
# Using ModelZoo models on your own datasets
-Animal behavior has to be analyzed with painstaking accuracy. Therefore, animal pose estimation has been an important tool to study animal behavior precisely. +
Animal behavior has to be analyzed with painstaking accuracy. Therefore, animal pose estimation has been
+an important tool to study animal behavior precisely.
-Beside providing an open source toolbox for researchers to develop customized deep neural networks for markerless pose estimation, we at DeepLabCut also aim to build robust, generalizable models. Part of this effort is via the [DeeplabCut ModelZoo](http://www.mackenziemathislab.org/dlc-modelzoo).
+Beside providing an open source toolbox for researchers to develop customized deep neural networks for markerless pose
+estimation, we at DeepLabCut also aim to build robust, generalizable models. Part of this effort is via the
+[DeeplabCut ModelZoo](http://modelzoo.deeplabcut.org/).
-The Zoo hosts user-contributed and #teamDLC developed models that are trained on specific animals and scenarios. You can analyze your videos directly with these models without training. The models have strong zero-shot performance on unseen out-of-domain data which can be further improved via pseudo-labeling. Please check the first [ModelZoo manuscript](https://arxiv.org/abs/2203.07436v1) for further details.
+The Zoo hosts user-contributed and DLC-team developed models that are trained on specific animals and scenarios. You can
+analyze your videos directly with these models without training. The models have strong zero-shot performance on unseen
+out-of-domain data which can be further improved via pseudo-labeling. Please check the first
+[ModelZoo manuscript](https://arxiv.org/abs/2203.07436v1) for further details.
-This recipe aims to show a usecase of the **mouse_pupil_vclose** and is contributed by 2022 DLC AI Resident [Neslihan Wittek](https://github.com/neslihanedes) 💜.
+This recipe aims to show a usecase of the **mouse_pupil_vclose** and is contributed by 2022 DLC AI Resident
+[Neslihan Wittek](https://github.com/neslihanedes) 💜.
## `mouse_pupil_vclose` model
This model was contributed by Jim McBurney-Lin at University of California Riverside, USA.
-The model was trained on images of C57/B6J mice eyes, and also then augmented with mouse eye data from the Mathis Lab at EPFL.
+The model was trained on images of C57/B6J mice eyes, and also then augmented with mouse eye data from the Mathis Lab at
+EPFL.
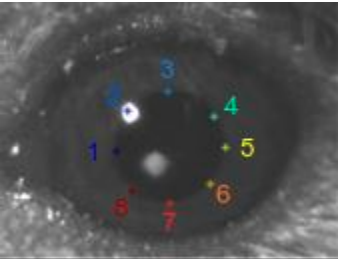 @@ -30,23 +38,37 @@ The model was trained on images of C57/B6J mice eyes, and also then augmented wi
| 8 | VLpupil | Ventral/left aspect of pupil |
-Since we would like to evaluate the models performance on out-of-domain data, we will analyze pigeon pupils. For more discussions and work on so-called out-of-domain data, see [Mathis, Biasi 2020](http://www.mackenziemathislab.org/horse10).
+Since we would like to evaluate the models performance on out-of-domain data, we will analyze pigeon pupils. For more
+discussions and work on so-called out-of-domain data, see
+[Mathis, Biasi 2020](https://paperswithcode.com/dataset/horse-10).
## Pigeon Pupil
-The eye pupil admits and regulates the amount of light entering the retina in order to enable image perception. Beside this curicial role, the pupil also reflects the state of the brain. The systemic behavior of the pupil has not been vastly studied in birds, although researchers from Max Planck Institute for Ornithology in Seewiesen have shed light on pupil behaviors in pigeons.
+The eye pupil admits and regulates the amount of light entering the retina in order to enable image perception. Beside
+this curicial role, the pupil also reflects the state of the brain. The systemic behavior of the pupil has not been
+vastly studied in birds, although researchers from
+Max Planck Institute for Ornithology in Seewiesen
+have shed light on pupil behaviors in pigeons.
-The pupils of male pigeons get smaller during courtship behavior. This is in contrast to mammals, for which the pupil size dilates in response to an increase in arousal. In addition, the pupil size of pigeons dilates during non-REM sleep, while they rapidly constrict during REM sleep. Examining these differences and the reason behind them, might be helpful to understand the pupillary behavior in general.
+The pupils of male pigeons get smaller during courtship behavior. This is in contrast to mammals, for which the pupil
+size dilates in response to an increase in arousal. In addition, the pupil size of pigeons dilates during non-REM sleep,
+while they rapidly constrict during REM sleep. Examining these differences and the reason behind them, might be helpful
+to understand the pupillary behavior in general.
-In light of these findings, we wanted to show whether the **mouse_pupil_vclose** model give us an accurate tracking performance for the pigeon pupil as well.
+In light of these findings, we wanted to show whether the **mouse_pupil_vclose** model give us an accurate tracking
+performance for the pigeon pupil as well.
### Jupyter & Google Colab Notebook
-DeepLabCut provides a Google Colab Notebook to analyze your video with a pretrained networks from the ModelZoo. No need for local installation of DeepLabCut!
+DeepLabCut provides a Google Colab Notebook to analyze your video with a pretrained networks from the ModelZoo. No need
+for local installation of DeepLabCut!
-Since we are interested in the accuracy of the **mouse_pupil_vclose** on pigeon pupil data, we will use a video which consists of 7 recordings of pigeon pupils.
+Since we are interested in the accuracy of the **mouse_pupil_vclose** on pigeon pupil data, we will use a video which
+consists of 7 recordings of pigeon pupils.
-Check ModelZoo Colab page and a video tutorial on how to use the ModelZoo on Google Colab.
+Check the
+[ModelZoo Colab page](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DLC_ModelZoo.ipynb)
+and a video tutorial on how to use the ModelZoo on Google Colab.
@@ -30,23 +38,37 @@ The model was trained on images of C57/B6J mice eyes, and also then augmented wi
| 8 | VLpupil | Ventral/left aspect of pupil |
-Since we would like to evaluate the models performance on out-of-domain data, we will analyze pigeon pupils. For more discussions and work on so-called out-of-domain data, see [Mathis, Biasi 2020](http://www.mackenziemathislab.org/horse10).
+Since we would like to evaluate the models performance on out-of-domain data, we will analyze pigeon pupils. For more
+discussions and work on so-called out-of-domain data, see
+[Mathis, Biasi 2020](https://paperswithcode.com/dataset/horse-10).
## Pigeon Pupil
-The eye pupil admits and regulates the amount of light entering the retina in order to enable image perception. Beside this curicial role, the pupil also reflects the state of the brain. The systemic behavior of the pupil has not been vastly studied in birds, although researchers from Max Planck Institute for Ornithology in Seewiesen have shed light on pupil behaviors in pigeons.
+The eye pupil admits and regulates the amount of light entering the retina in order to enable image perception. Beside
+this curicial role, the pupil also reflects the state of the brain. The systemic behavior of the pupil has not been
+vastly studied in birds, although researchers from
+Max Planck Institute for Ornithology in Seewiesen
+have shed light on pupil behaviors in pigeons.
-The pupils of male pigeons get smaller during courtship behavior. This is in contrast to mammals, for which the pupil size dilates in response to an increase in arousal. In addition, the pupil size of pigeons dilates during non-REM sleep, while they rapidly constrict during REM sleep. Examining these differences and the reason behind them, might be helpful to understand the pupillary behavior in general.
+The pupils of male pigeons get smaller during courtship behavior. This is in contrast to mammals, for which the pupil
+size dilates in response to an increase in arousal. In addition, the pupil size of pigeons dilates during non-REM sleep,
+while they rapidly constrict during REM sleep. Examining these differences and the reason behind them, might be helpful
+to understand the pupillary behavior in general.
-In light of these findings, we wanted to show whether the **mouse_pupil_vclose** model give us an accurate tracking performance for the pigeon pupil as well.
+In light of these findings, we wanted to show whether the **mouse_pupil_vclose** model give us an accurate tracking
+performance for the pigeon pupil as well.
### Jupyter & Google Colab Notebook
-DeepLabCut provides a Google Colab Notebook to analyze your video with a pretrained networks from the ModelZoo. No need for local installation of DeepLabCut!
+DeepLabCut provides a Google Colab Notebook to analyze your video with a pretrained networks from the ModelZoo. No need
+for local installation of DeepLabCut!
-Since we are interested in the accuracy of the **mouse_pupil_vclose** on pigeon pupil data, we will use a video which consists of 7 recordings of pigeon pupils.
+Since we are interested in the accuracy of the **mouse_pupil_vclose** on pigeon pupil data, we will use a video which
+consists of 7 recordings of pigeon pupils.
-Check ModelZoo Colab page and a video tutorial on how to use the ModelZoo on Google Colab.
+Check the
+[ModelZoo Colab page](https://github.com/DeepLabCut/DeepLabCut/blob/main/examples/COLAB/COLAB_DLC_ModelZoo.ipynb)
+and a video tutorial on how to use the ModelZoo on Google Colab.
 @@ -63,35 +85,36 @@ files.download("/content/file.zip")
### Analyze Videos at Your Local Machine
-DeepLabCut host models from the DeepLabCut ModelZoo Project.
+DeepLabCut host models from the [DeepLabCut ModelZoo Project](http://modelzoo.deeplabcut.org/).
The `create_pretrained_project` function will create a new project directory with the necessary sub-directories and a basic configuration file.
It will also initialize your project with a pre-trained model from the DeepLabCut ModelZoo.
The rest of the code should be run within your DeepLabCut environment.
-Check here for the instructions for the DeepLabCut installation.
+Check [here](how-to-install) for the instructions for the DeepLabCut installation.
+To initialize a new project directory with a pre-trained model from the DeepLabCut ModelZoo, run the code below.
+
+::::{warning}
+This method is currently implemented for Tensorflow only, Pytorch compatibility is coming soon.
+::::
```python
import deeplabcut
-```
-To initialize a new project directory with a pre-trained model from the DeepLabCut ModelZoo, run the code below.
-```python
deeplabcut.create_pretrained_project(
"projectname",
"experimenter",
- [r"path_for_the_videos"],
- model= "mouse_pupil_vclose",
- working_directory= r"project_directory",
- copy_videos= True,
- videotype= ".mp4 or .avi?",
- analyzevideo= True,
- filtered= True,
- createlabeledvideo= True,
- trainFraction= None
+ ["/home/max/Downloads/bunnies1.mp4"],
+ model="mouse_pupil_vclose",
+ working_directory="project_directory",
+ copy_videos=True,
+ analyzevideo=True,
+ filtered=True,
+ createlabeledvideo=True,
)
```
+
::::{important}
Your videos should be cropped around the eye for better model accuracy! 👁🐭
::::
@@ -100,13 +123,12 @@ Excitingly, 6 out of the 7 pigeon pupils were tracked nicely:
@@ -63,35 +85,36 @@ files.download("/content/file.zip")
### Analyze Videos at Your Local Machine
-DeepLabCut host models from the DeepLabCut ModelZoo Project.
+DeepLabCut host models from the [DeepLabCut ModelZoo Project](http://modelzoo.deeplabcut.org/).
The `create_pretrained_project` function will create a new project directory with the necessary sub-directories and a basic configuration file.
It will also initialize your project with a pre-trained model from the DeepLabCut ModelZoo.
The rest of the code should be run within your DeepLabCut environment.
-Check here for the instructions for the DeepLabCut installation.
+Check [here](how-to-install) for the instructions for the DeepLabCut installation.
+To initialize a new project directory with a pre-trained model from the DeepLabCut ModelZoo, run the code below.
+
+::::{warning}
+This method is currently implemented for Tensorflow only, Pytorch compatibility is coming soon.
+::::
```python
import deeplabcut
-```
-To initialize a new project directory with a pre-trained model from the DeepLabCut ModelZoo, run the code below.
-```python
deeplabcut.create_pretrained_project(
"projectname",
"experimenter",
- [r"path_for_the_videos"],
- model= "mouse_pupil_vclose",
- working_directory= r"project_directory",
- copy_videos= True,
- videotype= ".mp4 or .avi?",
- analyzevideo= True,
- filtered= True,
- createlabeledvideo= True,
- trainFraction= None
+ ["/home/max/Downloads/bunnies1.mp4"],
+ model="mouse_pupil_vclose",
+ working_directory="project_directory",
+ copy_videos=True,
+ analyzevideo=True,
+ filtered=True,
+ createlabeledvideo=True,
)
```
+
::::{important}
Your videos should be cropped around the eye for better model accuracy! 👁🐭
::::
@@ -100,13 +123,12 @@ Excitingly, 6 out of the 7 pigeon pupils were tracked nicely:
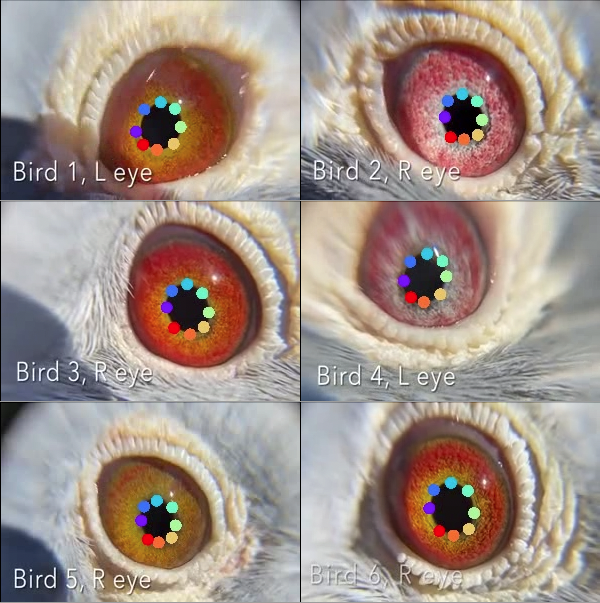 -
-When we further evaluate the model accuracy by checking the likelihood of tracked points, we see that the tracking is low confidience when the pigeons close their eyelid (which is of course expected, and can be leveraged to measure blinking 👁).
-
+When we further evaluate the model accuracy by checking the likelihood of tracked points, we see that the tracking is
+low confidience when the pigeons close their eyelid (which is of course expected, and can be leveraged to measure
+blinking 👁).
-
-When we further evaluate the model accuracy by checking the likelihood of tracked points, we see that the tracking is low confidience when the pigeons close their eyelid (which is of course expected, and can be leveraged to measure blinking 👁).
-
+When we further evaluate the model accuracy by checking the likelihood of tracked points, we see that the tracking is
+low confidience when the pigeons close their eyelid (which is of course expected, and can be leveraged to measure
+blinking 👁).
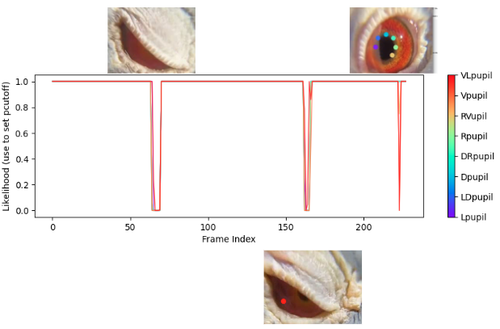 -
But you also might encounter larger problems than small tracking glitches:
-
But you also might encounter larger problems than small tracking glitches:
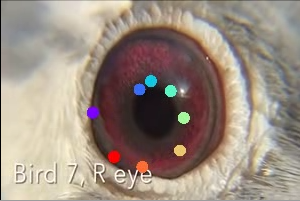 @@ -117,12 +139,26 @@ The more problems you encounter, the higher the number of frames you might want
You should also add the path of the video(s) into the `config.yaml` file, or run the following command to add the videos to your project:
```python
-deeplabcut.add_new_videos('/pathofproject/config.yaml', ['/pathofvideos/pigeon.mp4'], copy_videos=False, coords=None, extract_frames=False)
+deeplabcut.add_new_videos(
+ "/pathofproject/config.yaml",
+ ["/pathofvideos/pigeon.mp4"],
+ copy_videos=False,
+ coords=None,
+ extract_frames=False
+)
```
The `deeplabcut.extract_outlier_frames` function will check for outliers and ask your feedback on whether to extract these outliers frames.
```python
-deeplabcut.extract_outlier_frames('/pathofproject/config.yaml', ['/pathofvideos/pigeon.mp4'], automatic=True)
+deeplabcut.analyze_videos(
+ "/pathofproject/config.yaml",
+ ["/pathofvideos/pigeon.mp4"]
+)
+deeplabcut.extract_outlier_frames(
+ "/pathofproject/config.yaml",
+ ["/pathofvideos/pigeon.mp4"],
+ automatic=True
+)
```
The `deeplabcut.refine_labels` function starts the GUI which allows you to refine the outlier frames manually.
You should load the outlier frames directory and corresponding `.h5` file from the previous model.
@@ -130,9 +166,9 @@ It will ask you to define the `likelihood` threshold: labels under the threshold
After refining, you should combine these data with your previous model's data set and create a new training data set.
```python
-deeplabcut.refine_labels('/pathofproject/config.yaml')
-deeplabcut.merge_datasets('/pathofproject/config.yaml')
-deeplabcut.create_training_dataset('/pathofproject/config.yaml')
+deeplabcut.refine_labels("/pathofproject/config.yaml")
+deeplabcut.merge_datasets("/pathofproject/config.yaml")
+deeplabcut.create_training_dataset("/pathofproject/config.yaml")
```
Before starting the training of your model, there is one last step left: editing the `init_weights` parameter in your `pose_cfg.yaml` file.
Go to your project and check the latest snapshot (e.g., `snapshot-610000`) of your model in `dlc-models/train` directory.
@@ -142,7 +178,7 @@ Edit the value of the `init_weights` key in the `pose_cfg.yaml` file and start t
`init_weights: pathofyourproject\dlc-models\iteration-0\DLCFeb31-trainset95shuffle1\train\snapshot-610000`
```python
-deeplabcut.train_network('/pathofproject/config.yaml', shuffle=1, saveiters=25000)
+deeplabcut.train_network("/pathofproject/config.yaml", shuffle=1, saveiters=25000)
```
```{hint}
Check this video for model refining!
diff --git a/docs/recipes/nn.md b/docs/recipes/nn.md
index 75b44e315f..d35422fec9 100644
--- a/docs/recipes/nn.md
+++ b/docs/recipes/nn.md
@@ -1,3 +1,4 @@
+(tf-training-tips-and-tricks)=
# Model training tips & tricks
## Limiting a GPU's memory consumption
@@ -18,6 +19,7 @@ gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.25)
sess = tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
```
+(tf-custom-image-augmentation)=
## Using custom image augmentation
Image augmentation is the process of artificially expanding the training set
@@ -64,6 +66,7 @@ An in-depth tutorial on image augmentation and training hyperparameters can be f
The latest snapshot stored during training may not necessarily be the one that yields the highest performance. Therefore, you should analyze ALL snapshots, and select the best. Put 'all' in the snapshots section of the config.yaml to do this.
+(what-neural-network-should-i-use)=
## What neural network should I use? (Trade offs, speed performance, and considerations)
### With the release of even more network options, you now have to decide what to use! This additionally flexibility is hopefully helpful, but we want to give you some guidance on where to start.
diff --git a/docs/recipes/pose_cfg_file_breakdown.md b/docs/recipes/pose_cfg_file_breakdown.md
index 2e980edc25..2f79ac28d3 100644
--- a/docs/recipes/pose_cfg_file_breakdown.md
+++ b/docs/recipes/pose_cfg_file_breakdown.md
@@ -1,5 +1,10 @@
# The `pose_cfg.yaml` Guideline Handbook
+::::{warning}
+The following is specific to Tensorflow-based models. To read the equivalent explanations for Pytorch-based models,
+click [here](dlc3-pytorch-config)
+::::
+
👋 Hello! Mabuhay! Hola! This recipe was written by the [2023 DLC AI Residents](https://www.deeplabcutairesidency.org/)!
When you train, evaluate, and run inference with a neural network there are hyperparatmeters you must consider. While DLC attempts to set the "globally good for everyone" parameters, you might want to change them. Therefore, in this recipe we will review the pose config parameters related to neural network models' and the related data augmentation!
diff --git a/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md b/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md
index 31710c3a2d..4a7e47b856 100644
--- a/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md
+++ b/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md
@@ -133,7 +133,7 @@ This command installs DeepLabCut along with the dependencies required to build t
14. **🎉PR Approval:🎉** Once your PR is approved, the maintainers will merge it into the main repository. Your notebook will then be a part of the DeepLabCut Jupyter book! Yay!
-Remember to always check the [DLC CONTRIBUTING guidelines](https://github.com/DeepLabCut/DeepLabCut/blob/main/CONTRIBUTING.md).
+Remember to always check the [DLC contributing guidelines](https://github.com/DeepLabCut/DeepLabCut/blob/main/CONTRIBUTING.md).
## Wrap-Up 🎉
diff --git a/docs/roadmap.md b/docs/roadmap.md
index 905a95ae6f..b303754e37 100644
--- a/docs/roadmap.md
+++ b/docs/roadmap.md
@@ -2,7 +2,7 @@
## A development roadmap for DeepLabCut
-:loudspeaker: :hourglass_flowing_sand: :construction:
+📢 ⏳ 🚧
**General Enhancements:**
- [ ] DeepLabCut PyTorch & Model Zoo --> DLC 3.0 🔥
@@ -34,8 +34,8 @@
- [X] DeepLabCut-live! published in eLife
**DeepLabCut Model Zoo: a collection of pretrained models for plug-in-play DLC and community crowd-sourcing.**
-- [X] BETA release with 2.1.8b0: http://www.mousemotorlab.org/dlc-modelzoo
-- [X] full release with 2.1.8.1 http://www.mousemotorlab.org/dlc-modelzoo
+- [X] BETA release with 2.1.8b0: https://www.mackenziemathislab.org/deeplabcut
+- [X] full release with 2.1.8.1 https://www.mackenziemathislab.org/deeplabcut
- [X] Manuscript forthcoming! --> see arXiv https://arxiv.org/abs/2203.07436
- [X] new models added; horse, cheetah
- [X] TopView_Mouse model
diff --git a/docs/standardDeepLabCut_UserGuide.md b/docs/standardDeepLabCut_UserGuide.md
index 39188b3d48..7bbc1c71eb 100644
--- a/docs/standardDeepLabCut_UserGuide.md
+++ b/docs/standardDeepLabCut_UserGuide.md
@@ -1,7 +1,8 @@
(single-animal-userguide)=
# DeepLabCut User Guide (for single animal projects)
-This document covers single/standard DeepLabCut use. If you have a complicated multi-animal scenario (i.e., they look the same), then please see our [maDLC user guide](multi-animal-userguide).
+This document covers single/standard DeepLabCut use. If you have a complicated multi-animal scenario (i.e., they look
+the same), then please see our [maDLC user guide](multi-animal-userguide).
To get started, you can use the GUI, or the terminal. See below.
@@ -11,7 +12,12 @@ To get started, you can use the GUI, or the terminal. See below.
**GUI:**
-To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch "Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see Install docs!). Next, launch your conda env (i.e., for example `conda activate DEEPLABCUT`). Then, simply run ``python -m deeplabcut``. The below functions are available to you in an easy-to-use graphical user interface. While most functionality is available, advanced users might want the additional flexibility that command line interface offers. Read more below.
+To begin, navigate to Anaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch
+"Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see
+[install docs](how-to-install)!). Next, launch your conda env (i.e., for example `conda activate DEEPLABCUT`). Then,
+simply run `python -m deeplabcut`. The below functions are available to you in an easy-to-use graphical user interface.
+While most functionality is available, advanced users might want the additional flexibility that command line interface
+offers. Read more below.
```{Hint}
🚨 If you use Windows, please always open the terminal with administrator privileges! Right click, and "run as administrator".
```
@@ -20,11 +26,19 @@ To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admi
@@ -117,12 +139,26 @@ The more problems you encounter, the higher the number of frames you might want
You should also add the path of the video(s) into the `config.yaml` file, or run the following command to add the videos to your project:
```python
-deeplabcut.add_new_videos('/pathofproject/config.yaml', ['/pathofvideos/pigeon.mp4'], copy_videos=False, coords=None, extract_frames=False)
+deeplabcut.add_new_videos(
+ "/pathofproject/config.yaml",
+ ["/pathofvideos/pigeon.mp4"],
+ copy_videos=False,
+ coords=None,
+ extract_frames=False
+)
```
The `deeplabcut.extract_outlier_frames` function will check for outliers and ask your feedback on whether to extract these outliers frames.
```python
-deeplabcut.extract_outlier_frames('/pathofproject/config.yaml', ['/pathofvideos/pigeon.mp4'], automatic=True)
+deeplabcut.analyze_videos(
+ "/pathofproject/config.yaml",
+ ["/pathofvideos/pigeon.mp4"]
+)
+deeplabcut.extract_outlier_frames(
+ "/pathofproject/config.yaml",
+ ["/pathofvideos/pigeon.mp4"],
+ automatic=True
+)
```
The `deeplabcut.refine_labels` function starts the GUI which allows you to refine the outlier frames manually.
You should load the outlier frames directory and corresponding `.h5` file from the previous model.
@@ -130,9 +166,9 @@ It will ask you to define the `likelihood` threshold: labels under the threshold
After refining, you should combine these data with your previous model's data set and create a new training data set.
```python
-deeplabcut.refine_labels('/pathofproject/config.yaml')
-deeplabcut.merge_datasets('/pathofproject/config.yaml')
-deeplabcut.create_training_dataset('/pathofproject/config.yaml')
+deeplabcut.refine_labels("/pathofproject/config.yaml")
+deeplabcut.merge_datasets("/pathofproject/config.yaml")
+deeplabcut.create_training_dataset("/pathofproject/config.yaml")
```
Before starting the training of your model, there is one last step left: editing the `init_weights` parameter in your `pose_cfg.yaml` file.
Go to your project and check the latest snapshot (e.g., `snapshot-610000`) of your model in `dlc-models/train` directory.
@@ -142,7 +178,7 @@ Edit the value of the `init_weights` key in the `pose_cfg.yaml` file and start t
`init_weights: pathofyourproject\dlc-models\iteration-0\DLCFeb31-trainset95shuffle1\train\snapshot-610000`
```python
-deeplabcut.train_network('/pathofproject/config.yaml', shuffle=1, saveiters=25000)
+deeplabcut.train_network("/pathofproject/config.yaml", shuffle=1, saveiters=25000)
```
```{hint}
Check this video for model refining!
diff --git a/docs/recipes/nn.md b/docs/recipes/nn.md
index 75b44e315f..d35422fec9 100644
--- a/docs/recipes/nn.md
+++ b/docs/recipes/nn.md
@@ -1,3 +1,4 @@
+(tf-training-tips-and-tricks)=
# Model training tips & tricks
## Limiting a GPU's memory consumption
@@ -18,6 +19,7 @@ gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.25)
sess = tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
```
+(tf-custom-image-augmentation)=
## Using custom image augmentation
Image augmentation is the process of artificially expanding the training set
@@ -64,6 +66,7 @@ An in-depth tutorial on image augmentation and training hyperparameters can be f
The latest snapshot stored during training may not necessarily be the one that yields the highest performance. Therefore, you should analyze ALL snapshots, and select the best. Put 'all' in the snapshots section of the config.yaml to do this.
+(what-neural-network-should-i-use)=
## What neural network should I use? (Trade offs, speed performance, and considerations)
### With the release of even more network options, you now have to decide what to use! This additionally flexibility is hopefully helpful, but we want to give you some guidance on where to start.
diff --git a/docs/recipes/pose_cfg_file_breakdown.md b/docs/recipes/pose_cfg_file_breakdown.md
index 2e980edc25..2f79ac28d3 100644
--- a/docs/recipes/pose_cfg_file_breakdown.md
+++ b/docs/recipes/pose_cfg_file_breakdown.md
@@ -1,5 +1,10 @@
# The `pose_cfg.yaml` Guideline Handbook
+::::{warning}
+The following is specific to Tensorflow-based models. To read the equivalent explanations for Pytorch-based models,
+click [here](dlc3-pytorch-config)
+::::
+
👋 Hello! Mabuhay! Hola! This recipe was written by the [2023 DLC AI Residents](https://www.deeplabcutairesidency.org/)!
When you train, evaluate, and run inference with a neural network there are hyperparatmeters you must consider. While DLC attempts to set the "globally good for everyone" parameters, you might want to change them. Therefore, in this recipe we will review the pose config parameters related to neural network models' and the related data augmentation!
diff --git a/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md b/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md
index 31710c3a2d..4a7e47b856 100644
--- a/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md
+++ b/docs/recipes/publishing_notebooks_into_the_DLC_main_cookbook.md
@@ -133,7 +133,7 @@ This command installs DeepLabCut along with the dependencies required to build t
14. **🎉PR Approval:🎉** Once your PR is approved, the maintainers will merge it into the main repository. Your notebook will then be a part of the DeepLabCut Jupyter book! Yay!
-Remember to always check the [DLC CONTRIBUTING guidelines](https://github.com/DeepLabCut/DeepLabCut/blob/main/CONTRIBUTING.md).
+Remember to always check the [DLC contributing guidelines](https://github.com/DeepLabCut/DeepLabCut/blob/main/CONTRIBUTING.md).
## Wrap-Up 🎉
diff --git a/docs/roadmap.md b/docs/roadmap.md
index 905a95ae6f..b303754e37 100644
--- a/docs/roadmap.md
+++ b/docs/roadmap.md
@@ -2,7 +2,7 @@
## A development roadmap for DeepLabCut
-:loudspeaker: :hourglass_flowing_sand: :construction:
+📢 ⏳ 🚧
**General Enhancements:**
- [ ] DeepLabCut PyTorch & Model Zoo --> DLC 3.0 🔥
@@ -34,8 +34,8 @@
- [X] DeepLabCut-live! published in eLife
**DeepLabCut Model Zoo: a collection of pretrained models for plug-in-play DLC and community crowd-sourcing.**
-- [X] BETA release with 2.1.8b0: http://www.mousemotorlab.org/dlc-modelzoo
-- [X] full release with 2.1.8.1 http://www.mousemotorlab.org/dlc-modelzoo
+- [X] BETA release with 2.1.8b0: https://www.mackenziemathislab.org/deeplabcut
+- [X] full release with 2.1.8.1 https://www.mackenziemathislab.org/deeplabcut
- [X] Manuscript forthcoming! --> see arXiv https://arxiv.org/abs/2203.07436
- [X] new models added; horse, cheetah
- [X] TopView_Mouse model
diff --git a/docs/standardDeepLabCut_UserGuide.md b/docs/standardDeepLabCut_UserGuide.md
index 39188b3d48..7bbc1c71eb 100644
--- a/docs/standardDeepLabCut_UserGuide.md
+++ b/docs/standardDeepLabCut_UserGuide.md
@@ -1,7 +1,8 @@
(single-animal-userguide)=
# DeepLabCut User Guide (for single animal projects)
-This document covers single/standard DeepLabCut use. If you have a complicated multi-animal scenario (i.e., they look the same), then please see our [maDLC user guide](multi-animal-userguide).
+This document covers single/standard DeepLabCut use. If you have a complicated multi-animal scenario (i.e., they look
+the same), then please see our [maDLC user guide](multi-animal-userguide).
To get started, you can use the GUI, or the terminal. See below.
@@ -11,7 +12,12 @@ To get started, you can use the GUI, or the terminal. See below.
**GUI:**
-To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch "Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see Install docs!). Next, launch your conda env (i.e., for example `conda activate DEEPLABCUT`). Then, simply run ``python -m deeplabcut``. The below functions are available to you in an easy-to-use graphical user interface. While most functionality is available, advanced users might want the additional flexibility that command line interface offers. Read more below.
+To begin, navigate to Anaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch
+"Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see
+[install docs](how-to-install)!). Next, launch your conda env (i.e., for example `conda activate DEEPLABCUT`). Then,
+simply run `python -m deeplabcut`. The below functions are available to you in an easy-to-use graphical user interface.
+While most functionality is available, advanced users might want the additional flexibility that command line interface
+offers. Read more below.
```{Hint}
🚨 If you use Windows, please always open the terminal with administrator privileges! Right click, and "run as administrator".
```
@@ -20,11 +26,19 @@ To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admi
 -As a reminder, the core functions are described in our [Nature Protocols](https://www.nature.com/articles/s41596-019-0176-0) paper (published at the time of 2.0.6). Additional functions and features are continually added to the package. Thus, we recommend you read over the protocol and then please look at the following documentation and the doctrings. Thanks for using DeepLabCut!
+As a reminder, the core functions are described in our
+[Nature Protocols paper](https://www.nature.com/articles/s41596-019-0176-0) (published at the time of 2.0.6).
+Additional functions and features are continually added to the package. Thus, we recommend you read over the protocol
+and then please look at the following documentation and the doctrings. Thanks for using DeepLabCut!
## DeepLabCut in the Terminal/Command line interface:
-To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch "Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see Install docs!). Next, launch your conda env (i.e., for example `conda activate DEEPLABCUT`) and then type `ipython`. Then type `import deeplabcut`.
+To begin, navigate to Anaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch
+"Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see Install docs!). Next,
+launch your conda env (i.e., for example `conda activate DEEPLABCUT`) and then type `ipython`. Then type:
+```python
+import deeplabcut
+```
```{Hint}
🚨 If you use Windows, please always open the terminal with administrator privileges! Right click, and "run as administrator".
@@ -32,44 +46,87 @@ To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admi
### (A) Create a New Project
-The function **create\_new\_project** creates a new project directory, required subdirectories, and a basic project configuration file. Each project is identified by the name of the project (e.g. Reaching), name of the experimenter (e.g. YourName), as well as the date at creation.
+The function `create_new_project` creates a new project directory, required subdirectories, and a basic project
+configuration file. Each project is identified by the name of the project (e.g. Reaching), name of the experimenter
+(e.g. YourName), as well as the date at creation.
-Thus, this function requires the user to input the name of the project, the name of the experimenter, and the full path of the videos that are (initially) used to create the training dataset.
+Thus, this function requires the user to input the name of the project, the name of the experimenter, and the full
+path of the videos that are (initially) used to create the training dataset.
-Optional arguments specify the working directory, where the project directory will be created, and if the user wants to copy the videos (to the project directory). If the optional argument working\_directory is unspecified, the project directory is created in the current working directory, and if copy\_videos is unspecified symbolic links for the videos are created in the videos directory. Each symbolic link creates a reference to a video and thus eliminates the need to copy the entire video to the video directory (if the videos remain at the original location).
+Optional arguments specify the working directory, where the project directory will be created, and if the user wants
+to copy the videos (to the project directory). If the optional argument `working_directory` is unspecified, the
+project directory is created in the current working directory, and if `copy_videos` is unspecified symbolic links
+for the videos are created in the videos directory. Each symbolic link creates a reference to a video and thus
+eliminates the need to copy the entire video to the video directory (if the videos remain at the original location).
```python
-deeplabcut.create_new_project('Name of the project', 'Name of the experimenter', ['Full path of video 1', 'Full path of video2', 'Full path of video3'], working_directory='Full path of the working directory', copy_videos=True/False, multianimal=True/False)
+deeplabcut.create_new_project(
+ "Name of the project",
+ "Name of the experimenter",
+ ["Full path of video 1", "Full path of video2", "Full path of video3"],
+ working_directory="Full path of the working directory",
+ copy_videos=True/False,
+ multianimal=False
+)
```
**Important path formatting note**
-Windows users, you must input paths as: ``r'C:\Users\computername\Videos\reachingvideo1.avi' `` or
-
-`` 'C:\\Users\\computername\\Videos\\reachingvideo1.avi'``
-
- (TIP: you can also place ``config_path`` in front of ``deeplabcut.create_new_project`` to create a variable that holds the path to the config.yaml file, i.e. ``config_path=deeplabcut.create_new_project(...)``)
-
-
-This set of arguments will create a project directory with the name **Name of the project+name of the experimenter+date of creation of the project** in the **Working directory** and creates the symbolic links to videos in the **videos** directory. The project directory will have subdirectories: **dlc-models**, **labeled-data**, **training-datasets**, and **videos**. All the outputs generated during the course of a project will be stored in one of these subdirectories, thus allowing each project to be curated in separation from other projects. The purpose of the subdirectories is as follows:
-
-**dlc-models:** This directory contains the subdirectories *test* and *train*, each of which holds the meta information with regard to the parameters of the feature detectors in configuration files. The configuration files are YAML files, a common human-readable data serialization language. These files can be opened and edited with standard text editors. The subdirectory *train* will store checkpoints (called snapshots in TensorFlow) during training of the model. These snapshots allow the user to reload the trained model without re-training it, or to pick-up training from a particular saved checkpoint, in case the training was interrupted.
-
-**labeled-data:** This directory will store the frames used to create the training dataset. Frames from different videos are stored in separate subdirectories. Each frame has a filename related to the temporal index within the corresponding video, which allows the user to trace every frame back to its origin.
-
-**training-datasets:** This directory will contain the training dataset used to train the network and metadata, which contains information about how the training dataset was created.
-
-**videos:** Directory of video links or videos. When **copy\_videos** is set to ``False``, this directory contains symbolic links to the videos. If it is set to ``True`` then the videos will be copied to this directory. The default is ``False``. Additionally, if the user wants to add new videos to the project at any stage, the function **add\_new\_videos** can be used. This will update the list of videos in the project's configuration file.
+Windows users, you must input paths as: `r'C:\Users\computername\Videos\reachingvideo1.avi'` or
+` 'C:\\Users\\computername\\Videos\\reachingvideo1.avi'`
+
+TIP: you can also place `config_path` in front of `deeplabcut.create_new_project` to create a variable that holds
+the path to the config.yaml file, i.e. `config_path=deeplabcut.create_new_project(...)`
+
+This set of arguments will create a project directory with the name
+**
-As a reminder, the core functions are described in our [Nature Protocols](https://www.nature.com/articles/s41596-019-0176-0) paper (published at the time of 2.0.6). Additional functions and features are continually added to the package. Thus, we recommend you read over the protocol and then please look at the following documentation and the doctrings. Thanks for using DeepLabCut!
+As a reminder, the core functions are described in our
+[Nature Protocols paper](https://www.nature.com/articles/s41596-019-0176-0) (published at the time of 2.0.6).
+Additional functions and features are continually added to the package. Thus, we recommend you read over the protocol
+and then please look at the following documentation and the doctrings. Thanks for using DeepLabCut!
## DeepLabCut in the Terminal/Command line interface:
-To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch "Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see Install docs!). Next, launch your conda env (i.e., for example `conda activate DEEPLABCUT`) and then type `ipython`. Then type `import deeplabcut`.
+To begin, navigate to Anaconda Prompt Terminal and right-click to "open as admin "(Windows), or simply launch
+"Terminal" (unix/MacOS) on your computer. We assume you have DeepLabCut installed (if not, see Install docs!). Next,
+launch your conda env (i.e., for example `conda activate DEEPLABCUT`) and then type `ipython`. Then type:
+```python
+import deeplabcut
+```
```{Hint}
🚨 If you use Windows, please always open the terminal with administrator privileges! Right click, and "run as administrator".
@@ -32,44 +46,87 @@ To begin, navigate to Aanaconda Prompt Terminal and right-click to "open as admi
### (A) Create a New Project
-The function **create\_new\_project** creates a new project directory, required subdirectories, and a basic project configuration file. Each project is identified by the name of the project (e.g. Reaching), name of the experimenter (e.g. YourName), as well as the date at creation.
+The function `create_new_project` creates a new project directory, required subdirectories, and a basic project
+configuration file. Each project is identified by the name of the project (e.g. Reaching), name of the experimenter
+(e.g. YourName), as well as the date at creation.
-Thus, this function requires the user to input the name of the project, the name of the experimenter, and the full path of the videos that are (initially) used to create the training dataset.
+Thus, this function requires the user to input the name of the project, the name of the experimenter, and the full
+path of the videos that are (initially) used to create the training dataset.
-Optional arguments specify the working directory, where the project directory will be created, and if the user wants to copy the videos (to the project directory). If the optional argument working\_directory is unspecified, the project directory is created in the current working directory, and if copy\_videos is unspecified symbolic links for the videos are created in the videos directory. Each symbolic link creates a reference to a video and thus eliminates the need to copy the entire video to the video directory (if the videos remain at the original location).
+Optional arguments specify the working directory, where the project directory will be created, and if the user wants
+to copy the videos (to the project directory). If the optional argument `working_directory` is unspecified, the
+project directory is created in the current working directory, and if `copy_videos` is unspecified symbolic links
+for the videos are created in the videos directory. Each symbolic link creates a reference to a video and thus
+eliminates the need to copy the entire video to the video directory (if the videos remain at the original location).
```python
-deeplabcut.create_new_project('Name of the project', 'Name of the experimenter', ['Full path of video 1', 'Full path of video2', 'Full path of video3'], working_directory='Full path of the working directory', copy_videos=True/False, multianimal=True/False)
+deeplabcut.create_new_project(
+ "Name of the project",
+ "Name of the experimenter",
+ ["Full path of video 1", "Full path of video2", "Full path of video3"],
+ working_directory="Full path of the working directory",
+ copy_videos=True/False,
+ multianimal=False
+)
```
**Important path formatting note**
-Windows users, you must input paths as: ``r'C:\Users\computername\Videos\reachingvideo1.avi' `` or
-
-`` 'C:\\Users\\computername\\Videos\\reachingvideo1.avi'``
-
- (TIP: you can also place ``config_path`` in front of ``deeplabcut.create_new_project`` to create a variable that holds the path to the config.yaml file, i.e. ``config_path=deeplabcut.create_new_project(...)``)
-
-
-This set of arguments will create a project directory with the name **Name of the project+name of the experimenter+date of creation of the project** in the **Working directory** and creates the symbolic links to videos in the **videos** directory. The project directory will have subdirectories: **dlc-models**, **labeled-data**, **training-datasets**, and **videos**. All the outputs generated during the course of a project will be stored in one of these subdirectories, thus allowing each project to be curated in separation from other projects. The purpose of the subdirectories is as follows:
-
-**dlc-models:** This directory contains the subdirectories *test* and *train*, each of which holds the meta information with regard to the parameters of the feature detectors in configuration files. The configuration files are YAML files, a common human-readable data serialization language. These files can be opened and edited with standard text editors. The subdirectory *train* will store checkpoints (called snapshots in TensorFlow) during training of the model. These snapshots allow the user to reload the trained model without re-training it, or to pick-up training from a particular saved checkpoint, in case the training was interrupted.
-
-**labeled-data:** This directory will store the frames used to create the training dataset. Frames from different videos are stored in separate subdirectories. Each frame has a filename related to the temporal index within the corresponding video, which allows the user to trace every frame back to its origin.
-
-**training-datasets:** This directory will contain the training dataset used to train the network and metadata, which contains information about how the training dataset was created.
-
-**videos:** Directory of video links or videos. When **copy\_videos** is set to ``False``, this directory contains symbolic links to the videos. If it is set to ``True`` then the videos will be copied to this directory. The default is ``False``. Additionally, if the user wants to add new videos to the project at any stage, the function **add\_new\_videos** can be used. This will update the list of videos in the project's configuration file.
+Windows users, you must input paths as: `r'C:\Users\computername\Videos\reachingvideo1.avi'` or
+` 'C:\\Users\\computername\\Videos\\reachingvideo1.avi'`
+
+TIP: you can also place `config_path` in front of `deeplabcut.create_new_project` to create a variable that holds
+the path to the config.yaml file, i.e. `config_path=deeplabcut.create_new_project(...)`
+
+This set of arguments will create a project directory with the name
+**
 @@ -87,7 +144,11 @@ The ``create_new_project`` step writes the following parameters to the configura
@@ -87,7 +144,11 @@ The ``create_new_project`` step writes the following parameters to the configura
 -Next, open the **config.yaml** file, which was created during **create\_new\_project**. You can edit this file in any text editor. Familiarize yourself with the meaning of the parameters (Box 1). You can edit various parameters, in particular you **must add the list of *bodyparts* (or points of interest)** that you want to track. You can also set the *colormap* here that is used for all downstream steps (can also be edited at anytime), like labeling GUIs, videos, etc. Here any [matplotlib colormaps](https://matplotlib.org/tutorials/colors/colormaps.html) will do!
+Next, open the **config.yaml** file, which was created during **create\_new\_project**. You can edit this file in any
+text editor. Familiarize yourself with the meaning of the parameters (Box 1). You can edit various parameters, in
+particular you **must add the list of *bodyparts* (or points of interest)** that you want to track. You can also set the
+*colormap* here that is used for all downstream steps (can also be edited at anytime), like labeling GUIs, videos, etc.
+Here any [matplotlib colormaps](https://matplotlib.org/tutorials/colors/colormaps.html) will do!
Please DO NOT have spaces in the names of bodyparts.
**bodyparts:** are the bodyparts of each individual (in the above list).
@@ -95,26 +156,49 @@ Please DO NOT have spaces in the names of bodyparts.
### (C) Data Selection (extract frames)
-**CRITICAL:** A good training dataset should consist of a sufficient number of frames that capture the breadth of the behavior. This ideally implies to select the frames from different (behavioral) sessions, different lighting and different animals, if those vary substantially (to train an invariant, robust feature detector). Thus for creating a robust network that you can reuse in the laboratory, a good training dataset should reflect the diversity of the behavior with respect to postures, luminance conditions, background conditions, animal identities,etc. of the data that will be analyzed. For the simple lab behaviors comprising mouse reaching, open-field behavior and fly behavior, 100−200 frames gave good results [Mathis et al, 2018](https://www.nature.com/articles/s41593-018-0209-y). However, depending on the required accuracy, the nature of behavior, the video quality (e.g. motion blur, bad lighting) and the context, more or less frames might be necessary to create a good network. Ultimately, in order to scale up the analysis to large collections of videos with perhaps unexpected conditions, one can also refine the data set in an adaptive way (see refinement below).
-
-The function `extract_frames` extracts frames from all the videos in the project configuration file in order to create a training dataset. The extracted frames from all the videos are stored in a separate subdirectory named after the video file’s name under the ‘labeled-data’. This function also has various parameters that might be useful based on the user’s need.
+**CRITICAL:** A good training dataset should consist of a sufficient number of frames that capture the breadth of the
+behavior. This ideally implies to select the frames from different (behavioral) sessions, different lighting and
+different animals, if those vary substantially (to train an invariant, robust feature detector). Thus for creating a
+robust network that you can reuse in the laboratory, a good training dataset should reflect the diversity of the
+behavior with respect to postures, luminance conditions, background conditions, animal identities,etc. of the data that
+will be analyzed. For the simple lab behaviors comprising mouse reaching, open-field behavior and fly behavior, 100−200
+frames gave good results [Mathis et al, 2018](https://www.nature.com/articles/s41593-018-0209-y). However, depending on
+the required accuracy, the nature of behavior, the video quality (e.g. motion blur, bad lighting) and the context, more
+or less frames might be necessary to create a good network. Ultimately, in order to scale up the analysis to large
+collections of videos with perhaps unexpected conditions, one can also refine the data set in an adaptive way (see
+refinement below).
+
+The function `extract_frames` extracts frames from all the videos in the project configuration file in order to create
+a training dataset. The extracted frames from all the videos are stored in a separate subdirectory named after the video
+file’s name under the ‘labeled-data’. This function also has various parameters that might be useful based on the user’s
+need.
```python
-deeplabcut.extract_frames(config_path, mode='automatic/manual', algo='uniform/kmeans', userfeedback=False, crop=True/False)
+deeplabcut.extract_frames(
+ config_path,
+ mode="automatic/manual",
+ algo="uniform/kmeans",
+ crop=True/False,
+ userfeedback=False
+)
```
**CRITICAL POINT:** It is advisable to keep the frame size small, as large frames increase the training and
inference time. The cropping parameters for each video can be provided in the config.yaml file (and see below).
-When running the function extract_frames, if the parameter crop=True, then you will be asked to draw a box within the GUI (and this is written to the config.yaml file).
-
-`userfeedback` allows the user to check which videos they wish to extract frames from. In this way, if you added more videos to the config.yaml file it does not, by default, extract frames (again) from every video. If you wish to disable this question, set `userfeedback = True`.
-
-The provided function either selects frames from the videos that are randomly sampled from a uniform distribution (uniform), by clustering based on visual appearance (k-means), or by manual selection. Random
-selection of frames works best for behaviors where the postures vary across the whole video. However, some behaviors
-might be sparse, as in the case of reaching where the reach and pull are very fast and the mouse is not moving much
-between trials (thus, we have the default set to True, as this is best for most use-cases we encounter). In such a case, the function that allows selecting frames based on k-means derived quantization would
-be useful. If the user chooses to use k-means as a method to cluster the frames, then this function downsamples the
-video and clusters the frames using k-means, where each frame is treated as a vector. Frames from different clusters
-are then selected. This procedure makes sure that the frames look different. However, on large and long videos, this
-code is slow due to computational complexity.
+When running the function extract_frames, if the parameter crop=True, then you will be asked to draw a box within the
+GUI (and this is written to the config.yaml file).
+
+`userfeedback` allows the user to specify which videos they wish to extract frames from. When set to `"True"`, a dialog
+will be initiated, where the user is asked for each video if (additional/any) frames from this video should be
+extracted. Use this, e.g. if you have already labeled some folders and want to extract data for new videos.
+
+The provided function either selects frames from the videos that are randomly sampled from a uniform distribution
+(uniform), by clustering based on visual appearance (k-means), or by manual selection. Random uniform selection of
+frames works best for behaviors where the postures vary across the whole video. However, some behaviors might be sparse,
+as in the case of reaching where the reach and pull are very fast and the mouse is not moving much between trials. In
+such a case, the function that allows selecting frames based on k-means derived quantization would be useful. If the
+user chooses to use k-means as a method to cluster the frames, then this function downsamples the video and clusters the
+frames using k-means, where each frame is treated as a vector. Frames from different clusters are then selected. This
+procedure makes sure that the frames look different. However, on large and long videos, this code is slow due to
+computational complexity.
**CRITICAL POINT:** It is advisable to extract frames from a period of the video that contains interesting
behaviors, and not extract the frames across the whole video. This can be achieved by using the start and stop
@@ -122,13 +206,16 @@ parameters in the config.yaml file. Also, the user can change the number of fram
the numframes2extract in the config.yaml file.
However, picking frames is highly dependent on the data and the behavior being studied. Therefore, it is hard to
-provide all purpose code that extracts frames to create a good training dataset for every behavior and animal. If the user feels specific frames are lacking, they can extract hand selected frames of interest using the interactive GUI
+provide all purpose code that extracts frames to create a good training dataset for every behavior and animal. If the
+user feels specific frames are lacking, they can extract hand selected frames of interest using the interactive GUI
provided along with the toolbox. This can be launched by using:
```python
-deeplabcut.extract_frames(config_path, 'manual')
+deeplabcut.extract_frames(config_path, "manual")
```
The user can use the *Load Video* button to load one of the videos in the project configuration file, use the scroll
-bar to navigate across the video and *Grab a Frame* (or a range of frames, as of version 2.0.5) to extract the frame(s). The user can also look at the extracted frames and e.g. delete frames (from the directory) that are too similar before reloading the set and then manually annotating them.
+bar to navigate across the video and *Grab a Frame* (or a range of frames, as of version 2.0.5) to extract the frame(s).
+The user can also look at the extracted frames and e.g. delete frames (from the directory) that are too similar before
+reloading the set and then manually annotating them.
-Next, open the **config.yaml** file, which was created during **create\_new\_project**. You can edit this file in any text editor. Familiarize yourself with the meaning of the parameters (Box 1). You can edit various parameters, in particular you **must add the list of *bodyparts* (or points of interest)** that you want to track. You can also set the *colormap* here that is used for all downstream steps (can also be edited at anytime), like labeling GUIs, videos, etc. Here any [matplotlib colormaps](https://matplotlib.org/tutorials/colors/colormaps.html) will do!
+Next, open the **config.yaml** file, which was created during **create\_new\_project**. You can edit this file in any
+text editor. Familiarize yourself with the meaning of the parameters (Box 1). You can edit various parameters, in
+particular you **must add the list of *bodyparts* (or points of interest)** that you want to track. You can also set the
+*colormap* here that is used for all downstream steps (can also be edited at anytime), like labeling GUIs, videos, etc.
+Here any [matplotlib colormaps](https://matplotlib.org/tutorials/colors/colormaps.html) will do!
Please DO NOT have spaces in the names of bodyparts.
**bodyparts:** are the bodyparts of each individual (in the above list).
@@ -95,26 +156,49 @@ Please DO NOT have spaces in the names of bodyparts.
### (C) Data Selection (extract frames)
-**CRITICAL:** A good training dataset should consist of a sufficient number of frames that capture the breadth of the behavior. This ideally implies to select the frames from different (behavioral) sessions, different lighting and different animals, if those vary substantially (to train an invariant, robust feature detector). Thus for creating a robust network that you can reuse in the laboratory, a good training dataset should reflect the diversity of the behavior with respect to postures, luminance conditions, background conditions, animal identities,etc. of the data that will be analyzed. For the simple lab behaviors comprising mouse reaching, open-field behavior and fly behavior, 100−200 frames gave good results [Mathis et al, 2018](https://www.nature.com/articles/s41593-018-0209-y). However, depending on the required accuracy, the nature of behavior, the video quality (e.g. motion blur, bad lighting) and the context, more or less frames might be necessary to create a good network. Ultimately, in order to scale up the analysis to large collections of videos with perhaps unexpected conditions, one can also refine the data set in an adaptive way (see refinement below).
-
-The function `extract_frames` extracts frames from all the videos in the project configuration file in order to create a training dataset. The extracted frames from all the videos are stored in a separate subdirectory named after the video file’s name under the ‘labeled-data’. This function also has various parameters that might be useful based on the user’s need.
+**CRITICAL:** A good training dataset should consist of a sufficient number of frames that capture the breadth of the
+behavior. This ideally implies to select the frames from different (behavioral) sessions, different lighting and
+different animals, if those vary substantially (to train an invariant, robust feature detector). Thus for creating a
+robust network that you can reuse in the laboratory, a good training dataset should reflect the diversity of the
+behavior with respect to postures, luminance conditions, background conditions, animal identities,etc. of the data that
+will be analyzed. For the simple lab behaviors comprising mouse reaching, open-field behavior and fly behavior, 100−200
+frames gave good results [Mathis et al, 2018](https://www.nature.com/articles/s41593-018-0209-y). However, depending on
+the required accuracy, the nature of behavior, the video quality (e.g. motion blur, bad lighting) and the context, more
+or less frames might be necessary to create a good network. Ultimately, in order to scale up the analysis to large
+collections of videos with perhaps unexpected conditions, one can also refine the data set in an adaptive way (see
+refinement below).
+
+The function `extract_frames` extracts frames from all the videos in the project configuration file in order to create
+a training dataset. The extracted frames from all the videos are stored in a separate subdirectory named after the video
+file’s name under the ‘labeled-data’. This function also has various parameters that might be useful based on the user’s
+need.
```python
-deeplabcut.extract_frames(config_path, mode='automatic/manual', algo='uniform/kmeans', userfeedback=False, crop=True/False)
+deeplabcut.extract_frames(
+ config_path,
+ mode="automatic/manual",
+ algo="uniform/kmeans",
+ crop=True/False,
+ userfeedback=False
+)
```
**CRITICAL POINT:** It is advisable to keep the frame size small, as large frames increase the training and
inference time. The cropping parameters for each video can be provided in the config.yaml file (and see below).
-When running the function extract_frames, if the parameter crop=True, then you will be asked to draw a box within the GUI (and this is written to the config.yaml file).
-
-`userfeedback` allows the user to check which videos they wish to extract frames from. In this way, if you added more videos to the config.yaml file it does not, by default, extract frames (again) from every video. If you wish to disable this question, set `userfeedback = True`.
-
-The provided function either selects frames from the videos that are randomly sampled from a uniform distribution (uniform), by clustering based on visual appearance (k-means), or by manual selection. Random
-selection of frames works best for behaviors where the postures vary across the whole video. However, some behaviors
-might be sparse, as in the case of reaching where the reach and pull are very fast and the mouse is not moving much
-between trials (thus, we have the default set to True, as this is best for most use-cases we encounter). In such a case, the function that allows selecting frames based on k-means derived quantization would
-be useful. If the user chooses to use k-means as a method to cluster the frames, then this function downsamples the
-video and clusters the frames using k-means, where each frame is treated as a vector. Frames from different clusters
-are then selected. This procedure makes sure that the frames look different. However, on large and long videos, this
-code is slow due to computational complexity.
+When running the function extract_frames, if the parameter crop=True, then you will be asked to draw a box within the
+GUI (and this is written to the config.yaml file).
+
+`userfeedback` allows the user to specify which videos they wish to extract frames from. When set to `"True"`, a dialog
+will be initiated, where the user is asked for each video if (additional/any) frames from this video should be
+extracted. Use this, e.g. if you have already labeled some folders and want to extract data for new videos.
+
+The provided function either selects frames from the videos that are randomly sampled from a uniform distribution
+(uniform), by clustering based on visual appearance (k-means), or by manual selection. Random uniform selection of
+frames works best for behaviors where the postures vary across the whole video. However, some behaviors might be sparse,
+as in the case of reaching where the reach and pull are very fast and the mouse is not moving much between trials. In
+such a case, the function that allows selecting frames based on k-means derived quantization would be useful. If the
+user chooses to use k-means as a method to cluster the frames, then this function downsamples the video and clusters the
+frames using k-means, where each frame is treated as a vector. Frames from different clusters are then selected. This
+procedure makes sure that the frames look different. However, on large and long videos, this code is slow due to
+computational complexity.
**CRITICAL POINT:** It is advisable to extract frames from a period of the video that contains interesting
behaviors, and not extract the frames across the whole video. This can be achieved by using the start and stop
@@ -122,13 +206,16 @@ parameters in the config.yaml file. Also, the user can change the number of fram
the numframes2extract in the config.yaml file.
However, picking frames is highly dependent on the data and the behavior being studied. Therefore, it is hard to
-provide all purpose code that extracts frames to create a good training dataset for every behavior and animal. If the user feels specific frames are lacking, they can extract hand selected frames of interest using the interactive GUI
+provide all purpose code that extracts frames to create a good training dataset for every behavior and animal. If the
+user feels specific frames are lacking, they can extract hand selected frames of interest using the interactive GUI
provided along with the toolbox. This can be launched by using:
```python
-deeplabcut.extract_frames(config_path, 'manual')
+deeplabcut.extract_frames(config_path, "manual")
```
The user can use the *Load Video* button to load one of the videos in the project configuration file, use the scroll
-bar to navigate across the video and *Grab a Frame* (or a range of frames, as of version 2.0.5) to extract the frame(s). The user can also look at the extracted frames and e.g. delete frames (from the directory) that are too similar before reloading the set and then manually annotating them.
+bar to navigate across the video and *Grab a Frame* (or a range of frames, as of version 2.0.5) to extract the frame(s).
+The user can also look at the extracted frames and e.g. delete frames (from the directory) that are too similar before
+reloading the set and then manually annotating them.
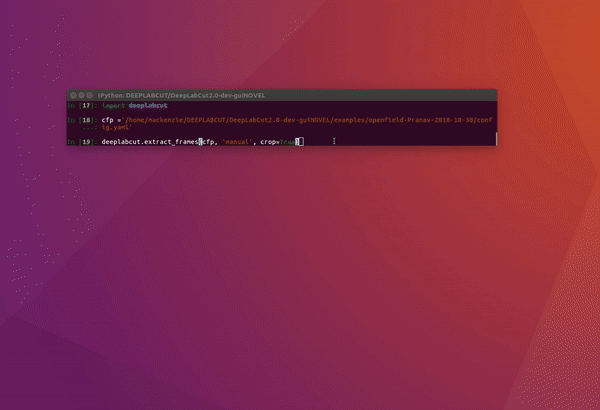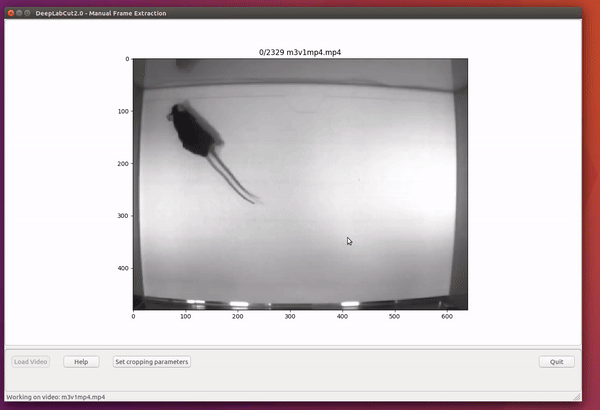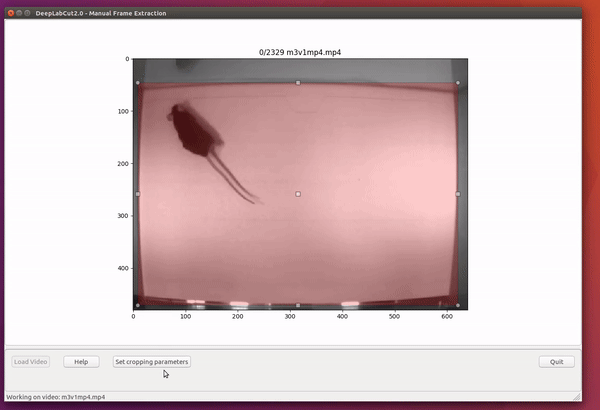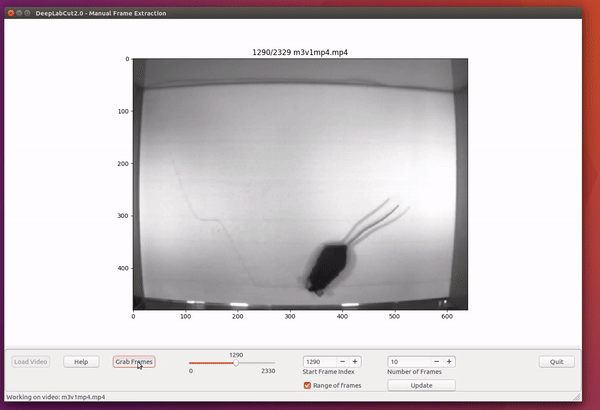 @@ -144,26 +231,25 @@ bar to navigate across the video and *Grab a Frame* (or a range of frames, as of
### (D) Label Frames
-The toolbox provides a function **label_frames** which helps the user to easily label all the extracted frames using
-an interactive graphical user interface (GUI). The user should have already named the body parts to label (points of
-interest) in the project’s configuration file by providing a list. The following command invokes the labeling toolbox.
+The toolbox provides a function **label_frames** which helps the user to easily label
+all the extracted frames using an interactive graphical user interface (GUI). The user
+should have already named the bodyparts to label (points of interest) in the
+project’s configuration file by providing a list. The following command invokes the
+napari-deeplabcut labelling GUI. Checkout the [napari-deeplabcut docs](napari-gui) for
+more information about the labelling workflow.
+
```python
deeplabcut.label_frames(config_path)
```
-The user needs to use the *Load Frames* button to select the directory which stores the extracted frames from one of
-the videos. Subsequently, the user can use one of the radio buttons (top right) to select a body part to label. RIGHT click to add the label. Left click to drag the label, if needed. If you label a part accidentally, you can use the middle button on your mouse to delete! If you cannot see a body part in the frame, skip over the label! Please see the ``HELP`` button for more user instructions. This auto-advances once you labeled the first body part. You can also advance to the next frame by clicking on the RIGHT arrow on your keyboard (and go to a previous frame with LEFT arrow).
-Each label will be plotted as a dot in a unique color.
-
-The user is free to move around the body part and once satisfied with its position, can select another radio button
-(in the top right) to switch to the respective body part (it otherwise auto-advances). The user can skip a body part if it is not visible. Once all the visible body parts are labeled, then the user can use ‘Next Frame’ to load the following frame. The user needs to save the labels after all the frames from one of the videos are labeled by clicking the save button at the bottom right. Saving the labels will create a labeled dataset for each video in a hierarchical data file format (HDF) in the
-subdirectory corresponding to the particular video in **labeled-data**. You can save at any intermediate step (even without closing the GUI, just hit save) and you return to labeling a dataset by reloading it!
**CRITICAL POINT:** It is advisable to **consistently label similar spots** (e.g., on a wrist that is very large, try
to label the same location). In general, invisible or occluded points should not be labeled by the user. They can
simply be skipped by not applying the label anywhere on the frame.
OPTIONAL: In the event of adding more labels to the existing labeled dataset, the user need to append the new
-labels to the bodyparts in the config.yaml file. Thereafter, the user can call the function **label_frames**. As of 2.0.5+: then a box will pop up and ask the user if they wish to display all parts, or only add in the new labels. Saving the labels after all the images are labelled will append the new labels to the existing labeled dataset.
+labels to the bodyparts in the config.yaml file. Thereafter, the user can call the function **label_frames**. As of
+2.0.5+: then a box will pop up and ask the user if they wish to display all parts, or only add in the new labels.
+Saving the labels after all the images are labelled will append the new labels to the existing labeled dataset.
HOT KEYS IN THE Labeling GUI (also see "help" in GUI):
```
@@ -173,7 +259,13 @@ Delete key: delete label.
```

-
+#### API Docs
+````{admonition} Click the button to see API Docs
+:class: dropdown
+```{eval-rst}
+.. include:: ./api/deeplabcut.label_frames.rst
+```
+````
### (E) Check Annotated Frames
@@ -184,7 +276,10 @@ is one of the most critical parts for creating the training dataset. The DeepLab
deeplabcut.check_labels(config_path, visualizeindividuals=True/False)
```
-For each video directory in labeled-data this function creates a subdirectory with **labeled** as a suffix. Those directories contain the frames plotted with the annotated body parts. The user can double check if the body parts are labeled correctly. If they are not correct, the user can reload the frames (i.e. `deeplabcut.label_frames`), move them around, and click save again.
+For each video directory in labeled-data this function creates a subdirectory with **labeled** as a suffix. Those
+directories contain the frames plotted with the annotated body parts. The user can double check if the body parts are
+labeled correctly. If they are not correct, the user can reload the frames (i.e. `deeplabcut.label_frames`), move them
+around, and click save again.
#### API Docs
````{admonition} Click the button to see API Docs
@@ -194,55 +289,81 @@ For each video directory in labeled-data this function creates a subdirectory wi
```
````
+(create-training-dataset)=
### (F) Create Training Dataset(s) and selection of your neural network
-**CRITICAL POINT:** Only run this step **where** you are going to train the network. If you label on your laptop but move your project folder to Google Colab or AWS, lab server, etc, then run the step below on that platform! If you labeled on a Windows machine but train on Linux, this is fine as of 2.0.4 onwards it will be done automatically (it saves file sets as both Linux and Windows for you).
+**CRITICAL POINT:** Only run this step **where** you are going to train the network. If you label on your laptop but
+move your project folder to Google Colab or AWS, lab server, etc, then run the step below on that platform! If you
+labeled on a Windows machine but train on Linux, this is fine as of 2.0.4 onwards it will be done automatically (it
+saves file sets as both Linux and Windows for you).
-- If you move your project folder, you must only change the `project_path` (which is done automatically) in the main config.yaml file - that's it - no need to change the video paths, etc! Your project is fully portable.
+- If you move your project folder, you must only change the `project_path` (which is done automatically) in the main
+config.yaml file - that's it - no need to change the video paths, etc! Your project is fully portable.
-- Be aware you select your neural network backbone at this stage. As of DLC3+ we support PyTorch (and TensorFlow, but this will be phased out).
+- Be aware you select your neural network backbone at this stage. As of DLC3+ we support PyTorch (and TensorFlow, but
+this will be phased out).
-**OVERVIEW:** This function combines the labeled datasets from all the videos and splits them to create train and test datasets. The training data will be used to train the network, while the test data set will be used for evaluating the network. The function **create_training_dataset** performs those steps.
+**OVERVIEW:** This function combines the labeled datasets from all the videos and splits them to create train and test
+datasets. The training data will be used to train the network, while the test data set will be used for evaluating the
+network.
```python
deeplabcut.create_training_dataset(config_path)
```
-- OPTIONAL: If the user wishes to benchmark the performance of the DeepLabCut, they can create multiple training datasets by specifying an integer value to the `num_shuffles`; see the docstring for more details.
+- OPTIONAL: If the user wishes to benchmark the performance of the DeepLabCut, they can create multiple training
+datasets by specifying an integer value to the `num_shuffles`; see the docstring for more details.
-within **dlc-models** called ``test`` and ``train``, and these each have a configuration file called pose_cfg.yaml.
-Specifically, the user can edit the **pose_cfg.yaml** within the **train** subdirectory before starting the training. These configuration files contain meta information with regard to the parameters of the feature detectors. Key parameters are listed in Box 2.
+The function creates a new shuffle(s) directory in the **dlc-models-pytorch** directory
+(**dlc-models** if using Tensorflow), in the current "iteration" directory.
+The `train` and `test` directories each have a configuration file
+(**pytorch_config.yaml** in **train** and **pose_cfg.yaml** in **test** for Pytorch models,
+**pose_cfg.yaml** in **train** and **test** for Tensorflow models).
+Specifically, the user can edit the **pytorch_config.yaml** (or **pose_cfg.yaml**) within the **train** subdirectory
+before starting the training. These configuration files contain meta information with regard to the parameters
+of the feature detectors. For more information about the **pytorch_config.yaml** file, see [here](dlc3-pytorch-config)
+(for TensorFlow-based models, see key parameters
+[here](https://github.com/DeepLabCut/DeepLabCut/blob/main/deeplabcut/pose_cfg.yaml)).
-**CRITICAL POINT:** At this step, for **create_training_dataset** you select the network you want to use, and any additional data augmentation (beyond our defaults). You can set ``net_type`` and ``augmenter_type`` when you call the function.
+**CRITICAL POINT:** At this step, for **create_training_dataset** you select the network you want to use, and any
+additional data augmentation (beyond our defaults). You can set `net_type`, `detector_type` (if using a detector)
+and `augmenter_type` when you call the function.
-- Networks: ImageNet pre-trained networks OR SuperAnimal pre-trained networks weights will be downloaded, as you select. You can decide to do transfer-learning (recommended) or "fine-tune" both the backbone and the decoder head. We suggest seeing our [dedicated documentation on models](https://deeplabcut.github.io/DeepLabCut/docs/pytorch/architectures.html) for more information.
+- Networks: ImageNet pre-trained networks OR SuperAnimal pre-trained networks weights will be downloaded, as you
+select. You can decide to do transfer-learning (recommended) or "fine-tune" both the backbone and the decoder head. We
+suggest seeing our [dedicated documentation on models](dlc3-architectures) for more information.
```{Hint}
-🚨 If they do not download (you will see this downloading in the terminal), then you may not have permission to do so - be sure to open your terminal "as an admin" (This is only something we have seen with some Windows users - see the **[docs for more help!](https://deeplabcut.github.io/DeepLabCut/docs/recipes/nn.html)**).
-```
-
-**DATA AUGMENTATION:** At this stage you can also decide what type of augmentation to use. The default loaders work well for most all tasks (as shown on www.deeplabcut.org), but there are many options, more data augmentation, intermediate supervision, etc. Please look at the [**pose_cfg.yaml**](https://github.com/DeepLabCut/DeepLabCut/blob/master/deeplabcut/pose_cfg.yaml) file for a full list of parameters **you might want to change before running this step.** There are several data loaders that can be used. For example, you can use the default loader (introduced and described in the Nature Protocols paper), [TensorPack](https://github.com/tensorpack/tensorpack) for data augmentation (currently this is easiest on Linux only), or [imgaug](https://imgaug.readthedocs.io/en/latest/). We recommend `imgaug`. You can set this by passing:``` deeplabcut.create_training_dataset(config_path, augmenter_type='imgaug') ```
-
-**For TensorFlow Models:** the differences of the loaders are as follows:
-- `imgaug`: a lot of augmentation possibilities, efficient code for target map creation & batch sizes >1 supported. You can set the parameters such as the `batch_size` in the `pose_cfg.yaml` file for the model you are training. This is the recommended DEFAULT!
-- `crop_scale`: our standard DLC 2.0 introduced in Nature Protocols variant (scaling, auto-crop augmentation)
-- `tensorpack`: a lot of augmentation possibilities, multi CPU support for fast processing, target maps are created less efficiently than in imgaug, does not allow batch size>1
-- `deterministic`: only useful for testing, freezes numpy seed; otherwise like default.
-
-**For PyTorch Models:**
-- #TODO: more information coming soon; in the meantime see the docstrings!
-
-
-**MODEL COMPARISON:** You can also test several models by creating the same test/train split for different networks. You can easily do this in the Project Manager GUI, which also lets you compare PyTorch and TensorFlow models.
-
-Please also consult the [following page on selecting models]( https://deeplabcut.github.io/DeepLabCut/docs/recipes/nn.html#what-neural-network-should-i-use-trade-offs-speed-performance-and-considerations)
-
- See Box 2 on how to specify **which network is loaded for training (including your own network, etc):**
-
-
@@ -144,26 +231,25 @@ bar to navigate across the video and *Grab a Frame* (or a range of frames, as of
### (D) Label Frames
-The toolbox provides a function **label_frames** which helps the user to easily label all the extracted frames using
-an interactive graphical user interface (GUI). The user should have already named the body parts to label (points of
-interest) in the project’s configuration file by providing a list. The following command invokes the labeling toolbox.
+The toolbox provides a function **label_frames** which helps the user to easily label
+all the extracted frames using an interactive graphical user interface (GUI). The user
+should have already named the bodyparts to label (points of interest) in the
+project’s configuration file by providing a list. The following command invokes the
+napari-deeplabcut labelling GUI. Checkout the [napari-deeplabcut docs](napari-gui) for
+more information about the labelling workflow.
+
```python
deeplabcut.label_frames(config_path)
```
-The user needs to use the *Load Frames* button to select the directory which stores the extracted frames from one of
-the videos. Subsequently, the user can use one of the radio buttons (top right) to select a body part to label. RIGHT click to add the label. Left click to drag the label, if needed. If you label a part accidentally, you can use the middle button on your mouse to delete! If you cannot see a body part in the frame, skip over the label! Please see the ``HELP`` button for more user instructions. This auto-advances once you labeled the first body part. You can also advance to the next frame by clicking on the RIGHT arrow on your keyboard (and go to a previous frame with LEFT arrow).
-Each label will be plotted as a dot in a unique color.
-
-The user is free to move around the body part and once satisfied with its position, can select another radio button
-(in the top right) to switch to the respective body part (it otherwise auto-advances). The user can skip a body part if it is not visible. Once all the visible body parts are labeled, then the user can use ‘Next Frame’ to load the following frame. The user needs to save the labels after all the frames from one of the videos are labeled by clicking the save button at the bottom right. Saving the labels will create a labeled dataset for each video in a hierarchical data file format (HDF) in the
-subdirectory corresponding to the particular video in **labeled-data**. You can save at any intermediate step (even without closing the GUI, just hit save) and you return to labeling a dataset by reloading it!
**CRITICAL POINT:** It is advisable to **consistently label similar spots** (e.g., on a wrist that is very large, try
to label the same location). In general, invisible or occluded points should not be labeled by the user. They can
simply be skipped by not applying the label anywhere on the frame.
OPTIONAL: In the event of adding more labels to the existing labeled dataset, the user need to append the new
-labels to the bodyparts in the config.yaml file. Thereafter, the user can call the function **label_frames**. As of 2.0.5+: then a box will pop up and ask the user if they wish to display all parts, or only add in the new labels. Saving the labels after all the images are labelled will append the new labels to the existing labeled dataset.
+labels to the bodyparts in the config.yaml file. Thereafter, the user can call the function **label_frames**. As of
+2.0.5+: then a box will pop up and ask the user if they wish to display all parts, or only add in the new labels.
+Saving the labels after all the images are labelled will append the new labels to the existing labeled dataset.
HOT KEYS IN THE Labeling GUI (also see "help" in GUI):
```
@@ -173,7 +259,13 @@ Delete key: delete label.
```

-
+#### API Docs
+````{admonition} Click the button to see API Docs
+:class: dropdown
+```{eval-rst}
+.. include:: ./api/deeplabcut.label_frames.rst
+```
+````
### (E) Check Annotated Frames
@@ -184,7 +276,10 @@ is one of the most critical parts for creating the training dataset. The DeepLab
deeplabcut.check_labels(config_path, visualizeindividuals=True/False)
```
-For each video directory in labeled-data this function creates a subdirectory with **labeled** as a suffix. Those directories contain the frames plotted with the annotated body parts. The user can double check if the body parts are labeled correctly. If they are not correct, the user can reload the frames (i.e. `deeplabcut.label_frames`), move them around, and click save again.
+For each video directory in labeled-data this function creates a subdirectory with **labeled** as a suffix. Those
+directories contain the frames plotted with the annotated body parts. The user can double check if the body parts are
+labeled correctly. If they are not correct, the user can reload the frames (i.e. `deeplabcut.label_frames`), move them
+around, and click save again.
#### API Docs
````{admonition} Click the button to see API Docs
@@ -194,55 +289,81 @@ For each video directory in labeled-data this function creates a subdirectory wi
```
````
+(create-training-dataset)=
### (F) Create Training Dataset(s) and selection of your neural network
-**CRITICAL POINT:** Only run this step **where** you are going to train the network. If you label on your laptop but move your project folder to Google Colab or AWS, lab server, etc, then run the step below on that platform! If you labeled on a Windows machine but train on Linux, this is fine as of 2.0.4 onwards it will be done automatically (it saves file sets as both Linux and Windows for you).
+**CRITICAL POINT:** Only run this step **where** you are going to train the network. If you label on your laptop but
+move your project folder to Google Colab or AWS, lab server, etc, then run the step below on that platform! If you
+labeled on a Windows machine but train on Linux, this is fine as of 2.0.4 onwards it will be done automatically (it
+saves file sets as both Linux and Windows for you).
-- If you move your project folder, you must only change the `project_path` (which is done automatically) in the main config.yaml file - that's it - no need to change the video paths, etc! Your project is fully portable.
+- If you move your project folder, you must only change the `project_path` (which is done automatically) in the main
+config.yaml file - that's it - no need to change the video paths, etc! Your project is fully portable.
-- Be aware you select your neural network backbone at this stage. As of DLC3+ we support PyTorch (and TensorFlow, but this will be phased out).
+- Be aware you select your neural network backbone at this stage. As of DLC3+ we support PyTorch (and TensorFlow, but
+this will be phased out).
-**OVERVIEW:** This function combines the labeled datasets from all the videos and splits them to create train and test datasets. The training data will be used to train the network, while the test data set will be used for evaluating the network. The function **create_training_dataset** performs those steps.
+**OVERVIEW:** This function combines the labeled datasets from all the videos and splits them to create train and test
+datasets. The training data will be used to train the network, while the test data set will be used for evaluating the
+network.
```python
deeplabcut.create_training_dataset(config_path)
```
-- OPTIONAL: If the user wishes to benchmark the performance of the DeepLabCut, they can create multiple training datasets by specifying an integer value to the `num_shuffles`; see the docstring for more details.
+- OPTIONAL: If the user wishes to benchmark the performance of the DeepLabCut, they can create multiple training
+datasets by specifying an integer value to the `num_shuffles`; see the docstring for more details.
-within **dlc-models** called ``test`` and ``train``, and these each have a configuration file called pose_cfg.yaml.
-Specifically, the user can edit the **pose_cfg.yaml** within the **train** subdirectory before starting the training. These configuration files contain meta information with regard to the parameters of the feature detectors. Key parameters are listed in Box 2.
+The function creates a new shuffle(s) directory in the **dlc-models-pytorch** directory
+(**dlc-models** if using Tensorflow), in the current "iteration" directory.
+The `train` and `test` directories each have a configuration file
+(**pytorch_config.yaml** in **train** and **pose_cfg.yaml** in **test** for Pytorch models,
+**pose_cfg.yaml** in **train** and **test** for Tensorflow models).
+Specifically, the user can edit the **pytorch_config.yaml** (or **pose_cfg.yaml**) within the **train** subdirectory
+before starting the training. These configuration files contain meta information with regard to the parameters
+of the feature detectors. For more information about the **pytorch_config.yaml** file, see [here](dlc3-pytorch-config)
+(for TensorFlow-based models, see key parameters
+[here](https://github.com/DeepLabCut/DeepLabCut/blob/main/deeplabcut/pose_cfg.yaml)).
-**CRITICAL POINT:** At this step, for **create_training_dataset** you select the network you want to use, and any additional data augmentation (beyond our defaults). You can set ``net_type`` and ``augmenter_type`` when you call the function.
+**CRITICAL POINT:** At this step, for **create_training_dataset** you select the network you want to use, and any
+additional data augmentation (beyond our defaults). You can set `net_type`, `detector_type` (if using a detector)
+and `augmenter_type` when you call the function.
-- Networks: ImageNet pre-trained networks OR SuperAnimal pre-trained networks weights will be downloaded, as you select. You can decide to do transfer-learning (recommended) or "fine-tune" both the backbone and the decoder head. We suggest seeing our [dedicated documentation on models](https://deeplabcut.github.io/DeepLabCut/docs/pytorch/architectures.html) for more information.
+- Networks: ImageNet pre-trained networks OR SuperAnimal pre-trained networks weights will be downloaded, as you
+select. You can decide to do transfer-learning (recommended) or "fine-tune" both the backbone and the decoder head. We
+suggest seeing our [dedicated documentation on models](dlc3-architectures) for more information.
```{Hint}
-🚨 If they do not download (you will see this downloading in the terminal), then you may not have permission to do so - be sure to open your terminal "as an admin" (This is only something we have seen with some Windows users - see the **[docs for more help!](https://deeplabcut.github.io/DeepLabCut/docs/recipes/nn.html)**).
-```
-
-**DATA AUGMENTATION:** At this stage you can also decide what type of augmentation to use. The default loaders work well for most all tasks (as shown on www.deeplabcut.org), but there are many options, more data augmentation, intermediate supervision, etc. Please look at the [**pose_cfg.yaml**](https://github.com/DeepLabCut/DeepLabCut/blob/master/deeplabcut/pose_cfg.yaml) file for a full list of parameters **you might want to change before running this step.** There are several data loaders that can be used. For example, you can use the default loader (introduced and described in the Nature Protocols paper), [TensorPack](https://github.com/tensorpack/tensorpack) for data augmentation (currently this is easiest on Linux only), or [imgaug](https://imgaug.readthedocs.io/en/latest/). We recommend `imgaug`. You can set this by passing:``` deeplabcut.create_training_dataset(config_path, augmenter_type='imgaug') ```
-
-**For TensorFlow Models:** the differences of the loaders are as follows:
-- `imgaug`: a lot of augmentation possibilities, efficient code for target map creation & batch sizes >1 supported. You can set the parameters such as the `batch_size` in the `pose_cfg.yaml` file for the model you are training. This is the recommended DEFAULT!
-- `crop_scale`: our standard DLC 2.0 introduced in Nature Protocols variant (scaling, auto-crop augmentation)
-- `tensorpack`: a lot of augmentation possibilities, multi CPU support for fast processing, target maps are created less efficiently than in imgaug, does not allow batch size>1
-- `deterministic`: only useful for testing, freezes numpy seed; otherwise like default.
-
-**For PyTorch Models:**
-- #TODO: more information coming soon; in the meantime see the docstrings!
-
-
-**MODEL COMPARISON:** You can also test several models by creating the same test/train split for different networks. You can easily do this in the Project Manager GUI, which also lets you compare PyTorch and TensorFlow models.
-
-Please also consult the [following page on selecting models]( https://deeplabcut.github.io/DeepLabCut/docs/recipes/nn.html#what-neural-network-should-i-use-trade-offs-speed-performance-and-considerations)
-
- See Box 2 on how to specify **which network is loaded for training (including your own network, etc):**
-
-
- -
-
 @@ -457,48 +653,85 @@ It creates a folder called ``plot-poses`` (in the directory of the video). The p
### (L) Create Labeled Videos:
Additionally, the toolbox provides a function to create labeled videos based on the extracted poses by plotting the
-labels on top of the frame and creating a video. There are two modes to create videos: FAST and SLOW (but higher quality!). If you want to create high-quality videos, please add ``save_frames=True``. One can use the command as follows to create multiple labeled videos:
+labels on top of the frame and creating a video. There are two modes to create videos: FAST and SLOW (but higher
+quality!). One can use the command as follows to create multiple labeled videos:
```python
-deeplabcut.create_labeled_video(config_path, ['fullpath/analysis/project/videos/reachingvideo1.avi','fullpath/analysis/project/videos/reachingvideo2.avi'], save_frames = True/False)
-```
- Optionally, if you want to use the filtered data for a video or directory of filtered videos pass ``filtered=True``, i.e.:
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/analysis/project/videos/reachingvideo1.avi",
+ "fullpath/analysis/project/videos/reachingvideo2.avi"],
+ save_frames = True/False
+)
+```
+ Optionally, if you want to use the filtered data for a video or directory of filtered videos pass `filtered=True`,
+ i.e.:
```python
-deeplabcut.create_labeled_video(config_path, ['fullpath/afolderofvideos'], videotype='.mp4', filtered=True)
-```
-You can also optionally add a skeleton to connect points and/or add a history of points for visualization. To set the "trailing points" you need to pass ``trailpoints``:
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ filtered=True
+)
+```
+You can also optionally add a skeleton to connect points and/or add a history of points for visualization. To set the
+"trailing points" you need to pass `trailpoints`:
```python
-deeplabcut.create_labeled_video(config_path, ['fullpath/afolderofvideos'], videotype='.mp4', trailpoints=10)
-```
-To draw a skeleton, you need to first define the pairs of connected nodes (in the ``config.yaml`` file) and set the skeleton color (in the ``config.yaml`` file). There is also a GUI to help you do this, use by calling `deeplabcut.SkeletonBuilder(config+path)`!
-
-Here is how the ``config.yaml`` additions/edits should look (for example, on the Openfield demo data we provide):
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ trailpoints=10
+)
+```
+To draw a skeleton, you need to first define the pairs of connected nodes (in the `config.yaml` file) and set the
+skeleton color (in the `config.yaml` file). There is also a GUI to help you do this, use by calling
+`deeplabcut.SkeletonBuilder(configpath)`!
+
+Here is how the `config.yaml` additions/edits should look (for example, on the Openfield demo data we provide):
```python
# Plotting configuration
-skeleton: [['snout', 'leftear'], ['snout', 'rightear'], ['leftear', 'tailbase'], ['leftear', 'rightear'], ['rightear','tailbase']]
+skeleton:
+ - ["snout", "leftear"]
+ - ["snout", "rightear"]
+ - ["leftear", "tailbase"]
+ - ["leftear", "rightear"]
+ - ["rightear", "tailbase"]
skeleton_color: white
pcutoff: 0.4
dotsize: 4
alphavalue: 0.5
colormap: jet
```
-Then pass ``draw_skeleton=True`` with the command:
+Then pass `draw_skeleton=True` with the command:
```python
-deeplabcut.create_labeled_video(config_path,['fullpath/afolderofvideos'], videotype='.mp4', draw_skeleton = True)
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ draw_skeleton=True
+)
```
-**NEW** as of 2.2b8: You can create a video with only the "dots" plotted, i.e., in the [style of Johansson](https://link.springer.com/article/10.1007/BF00309043), by passing `keypoints_only=True`:
+**NEW** as of 2.2b8: You can create a video with only the "dots" plotted, i.e., in the
+[style of Johansson](https://link.springer.com/article/10.1007/BF00309043), by passing `keypoints_only=True`:
```python
-deeplabcut.create_labeled_video(config_path,['fullpath/afolderofvideos'], videotype='.mp4', keypoints_only=True)
+deeplabcut.create_labeled_video(
+ config_path,["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ keypoints_only=True
+)
```
-**PRO TIP:** that the **best quality videos** are created when ``save_frames=True`` is passed. Therefore, when ``trailpoints`` and ``draw_skeleton`` are used, we **highly** recommend you also pass ``save_frames=True``!
+**PRO TIP:** that the **best quality videos** are created when `fastmode=False` is passed. Therefore, when
+`trailpoints` and `draw_skeleton` are used, we **highly** recommend you also pass `fastmode=False`!
@@ -457,48 +653,85 @@ It creates a folder called ``plot-poses`` (in the directory of the video). The p
### (L) Create Labeled Videos:
Additionally, the toolbox provides a function to create labeled videos based on the extracted poses by plotting the
-labels on top of the frame and creating a video. There are two modes to create videos: FAST and SLOW (but higher quality!). If you want to create high-quality videos, please add ``save_frames=True``. One can use the command as follows to create multiple labeled videos:
+labels on top of the frame and creating a video. There are two modes to create videos: FAST and SLOW (but higher
+quality!). One can use the command as follows to create multiple labeled videos:
```python
-deeplabcut.create_labeled_video(config_path, ['fullpath/analysis/project/videos/reachingvideo1.avi','fullpath/analysis/project/videos/reachingvideo2.avi'], save_frames = True/False)
-```
- Optionally, if you want to use the filtered data for a video or directory of filtered videos pass ``filtered=True``, i.e.:
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/analysis/project/videos/reachingvideo1.avi",
+ "fullpath/analysis/project/videos/reachingvideo2.avi"],
+ save_frames = True/False
+)
+```
+ Optionally, if you want to use the filtered data for a video or directory of filtered videos pass `filtered=True`,
+ i.e.:
```python
-deeplabcut.create_labeled_video(config_path, ['fullpath/afolderofvideos'], videotype='.mp4', filtered=True)
-```
-You can also optionally add a skeleton to connect points and/or add a history of points for visualization. To set the "trailing points" you need to pass ``trailpoints``:
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ filtered=True
+)
+```
+You can also optionally add a skeleton to connect points and/or add a history of points for visualization. To set the
+"trailing points" you need to pass `trailpoints`:
```python
-deeplabcut.create_labeled_video(config_path, ['fullpath/afolderofvideos'], videotype='.mp4', trailpoints=10)
-```
-To draw a skeleton, you need to first define the pairs of connected nodes (in the ``config.yaml`` file) and set the skeleton color (in the ``config.yaml`` file). There is also a GUI to help you do this, use by calling `deeplabcut.SkeletonBuilder(config+path)`!
-
-Here is how the ``config.yaml`` additions/edits should look (for example, on the Openfield demo data we provide):
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ trailpoints=10
+)
+```
+To draw a skeleton, you need to first define the pairs of connected nodes (in the `config.yaml` file) and set the
+skeleton color (in the `config.yaml` file). There is also a GUI to help you do this, use by calling
+`deeplabcut.SkeletonBuilder(configpath)`!
+
+Here is how the `config.yaml` additions/edits should look (for example, on the Openfield demo data we provide):
```python
# Plotting configuration
-skeleton: [['snout', 'leftear'], ['snout', 'rightear'], ['leftear', 'tailbase'], ['leftear', 'rightear'], ['rightear','tailbase']]
+skeleton:
+ - ["snout", "leftear"]
+ - ["snout", "rightear"]
+ - ["leftear", "tailbase"]
+ - ["leftear", "rightear"]
+ - ["rightear", "tailbase"]
skeleton_color: white
pcutoff: 0.4
dotsize: 4
alphavalue: 0.5
colormap: jet
```
-Then pass ``draw_skeleton=True`` with the command:
+Then pass `draw_skeleton=True` with the command:
```python
-deeplabcut.create_labeled_video(config_path,['fullpath/afolderofvideos'], videotype='.mp4', draw_skeleton = True)
+deeplabcut.create_labeled_video(
+ config_path,
+ ["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ draw_skeleton=True
+)
```
-**NEW** as of 2.2b8: You can create a video with only the "dots" plotted, i.e., in the [style of Johansson](https://link.springer.com/article/10.1007/BF00309043), by passing `keypoints_only=True`:
+**NEW** as of 2.2b8: You can create a video with only the "dots" plotted, i.e., in the
+[style of Johansson](https://link.springer.com/article/10.1007/BF00309043), by passing `keypoints_only=True`:
```python
-deeplabcut.create_labeled_video(config_path,['fullpath/afolderofvideos'], videotype='.mp4', keypoints_only=True)
+deeplabcut.create_labeled_video(
+ config_path,["fullpath/afolderofvideos"],
+ videotype=".mp4",
+ keypoints_only=True
+)
```
-**PRO TIP:** that the **best quality videos** are created when ``save_frames=True`` is passed. Therefore, when ``trailpoints`` and ``draw_skeleton`` are used, we **highly** recommend you also pass ``save_frames=True``!
+**PRO TIP:** that the **best quality videos** are created when `fastmode=False` is passed. Therefore, when
+`trailpoints` and `draw_skeleton` are used, we **highly** recommend you also pass `fastmode=False`!
